FHIR defines standardized resources, including Patient, Observation, Condition, Medication, Encounter, and DiagnosticReport that enable health data exchange across electronic health records, registries, analytics platforms, and artificial intelligence services. By representing data in FHIR format, diverse systems can reliably consume, transform, and act on health information without custom adapters or fragile point-to-point integrations. FHIR-based systems facilitate digital quality initiatives, including population health, quality measure reporting, and outcomes tracking, while accelerating data sharing and standardization across institutions.
However, much valuable clinical information remains locked in unstructured data including physician notes, radiology reports, discharge summaries, pathology narratives, and scanned documents. Symptoms, disease history, imaging interpretations, physical exam findings, social determinants, and follow-up plans are in many cases stored in unstructured text, not in structured fields. Reliance solely on structured electronic health record fields, billing codes, or manually entered data misses nuance, incomplete coding, and clinician reasoning, creating gaps in quality measure capture and limiting analytics potential. Manual abstraction to convert unstructured data into structured records is labor-intensive, slow, expensive, error-prone, and cannot scale with growing data volumes.
Why unstructured data prevents FHIR interoperability at scale
Real Chemistry’s pharma medical affairs integration demonstrates the unstructured data challenge. Their client’s medical affairs group needed to harmonize numerous unstructured datasets to build dashboards identifying areas of impact and opportunity. They leveraged Healthcare NLP’s medical entity extraction and sentiment analysis capabilities across proprietary social listening data and multiple client datasets, creating a unified view covering over 10 large and continuously evolving datasets.*
Without systematic extraction and normalization to standard terminologies, these datasets remained siloed, each using different vocabulary, coding conventions, and data structures. FHIR resource generation requires standardized terminology: conditions mapped to SNOMED CT, medications to RxNorm, laboratory tests to LOINC, and diagnoses to ICD-10. Unstructured text mentioning “MI” versus “myocardial infarction” versus “heart attack” must normalize to the same SNOMED CT concept before FHIR Condition resources can be reliably generated and exchanged.
Cancer registry building with large language models shows similar challenges. Teams evaluated relative performance of models for cancer disease response classification and sites of metastases identification, seeking low-cost, sustainable solutions for lean, low-resource healthcare data registry teams.* Registries require FHIR-compliant data submissions, yet cancer staging, biomarker results, treatment responses, and metastases locations are predominantly documented in unstructured pathology reports and clinical notes. Without automated extraction and terminology normalization, registry population remains prohibitively manual.
The six-layer architecture from unstructured text to FHIR resources
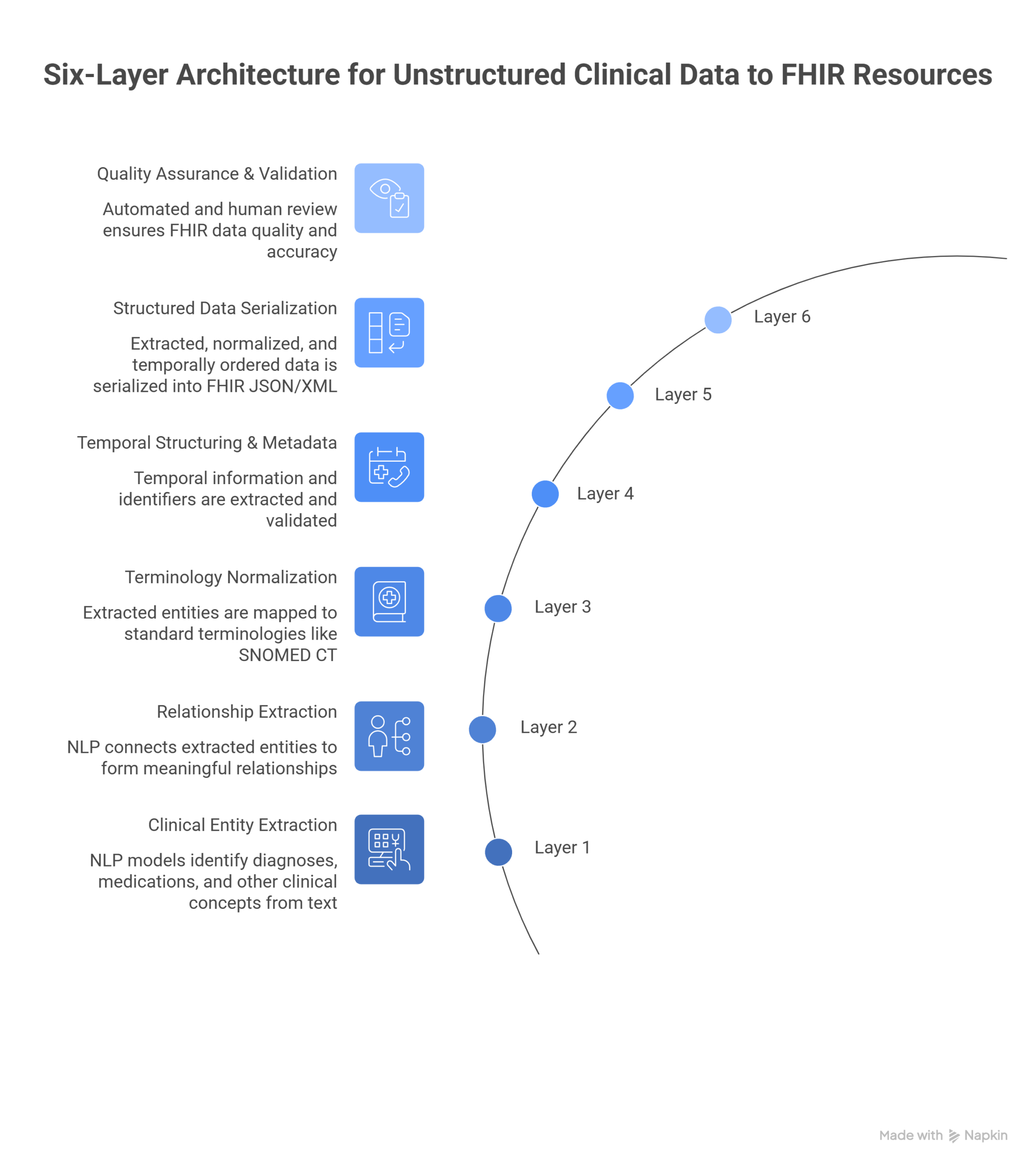
Healthcare NLP provides over 2,800 pre-trained clinical models extracting diagnoses, medications, procedures, laboratory tests, anatomical locations, and clinical findings from unstructured text. But extraction must capture clinical context and assertion detection distinguishing present conditions from absent, historical, or family history mentions. “No evidence of pneumonia” must not generate a FHIR Condition resource for pneumonia. “Family history of diabetes” requires different FHIR representation than “patient has diabetes.”
Layer 2: Relationship extraction connecting clinical concepts
FHIR resources include references linking related concepts: Medication resources reference Condition resources indicating treatment purpose, Observation resources link to Encounter resources establishing clinical context, Procedure resources connect to DiagnosticReport resources showing indications. Extracting these relationships from narrative text enables accurate FHIR resource generation. “Started metformin for diabetes” must create linked MedicationStatement and Condition resources, not isolated entities.
Layer 3: Terminology normalization to standard vocabularies
Healthcare NLP’s entity resolution capabilities map extracted entities to SNOMED CT, ICD-10, LOINC, and RxNorm. “Blood sugar” becomes LOINC code for glucose. “Heart attack” becomes SNOMED CT concept for myocardial infarction. “Tylenol” becomes RxNorm code for acetaminophen. This normalization enables semantic interoperability. As a result systems can recognize equivalent concepts even when documented differently.
Real Chemistry’s harmonization of over 10 datasets demonstrates why normalization matters for integration. Without mapping client-specific terminology, product naming conventions, and therapeutic area taxonomies to standard vocabularies, unified dashboards cannot aggregate information consistently across evolving datasets.
Layer 4: Temporal structuring and metadata extraction
FHIR resources require temporal information: onset dates for conditions, administration times for medications, result timestamps for observations. “Started on aspirin three months ago” must extract both the medication (aspirin/RxNorm code) and temporal relationship (three months prior to note date) to generate accurate MedicationStatement resources with onset datetime. Patient identifiers, encounter identifiers, and practitioner references must be extracted and validated to populate required FHIR resource fields.
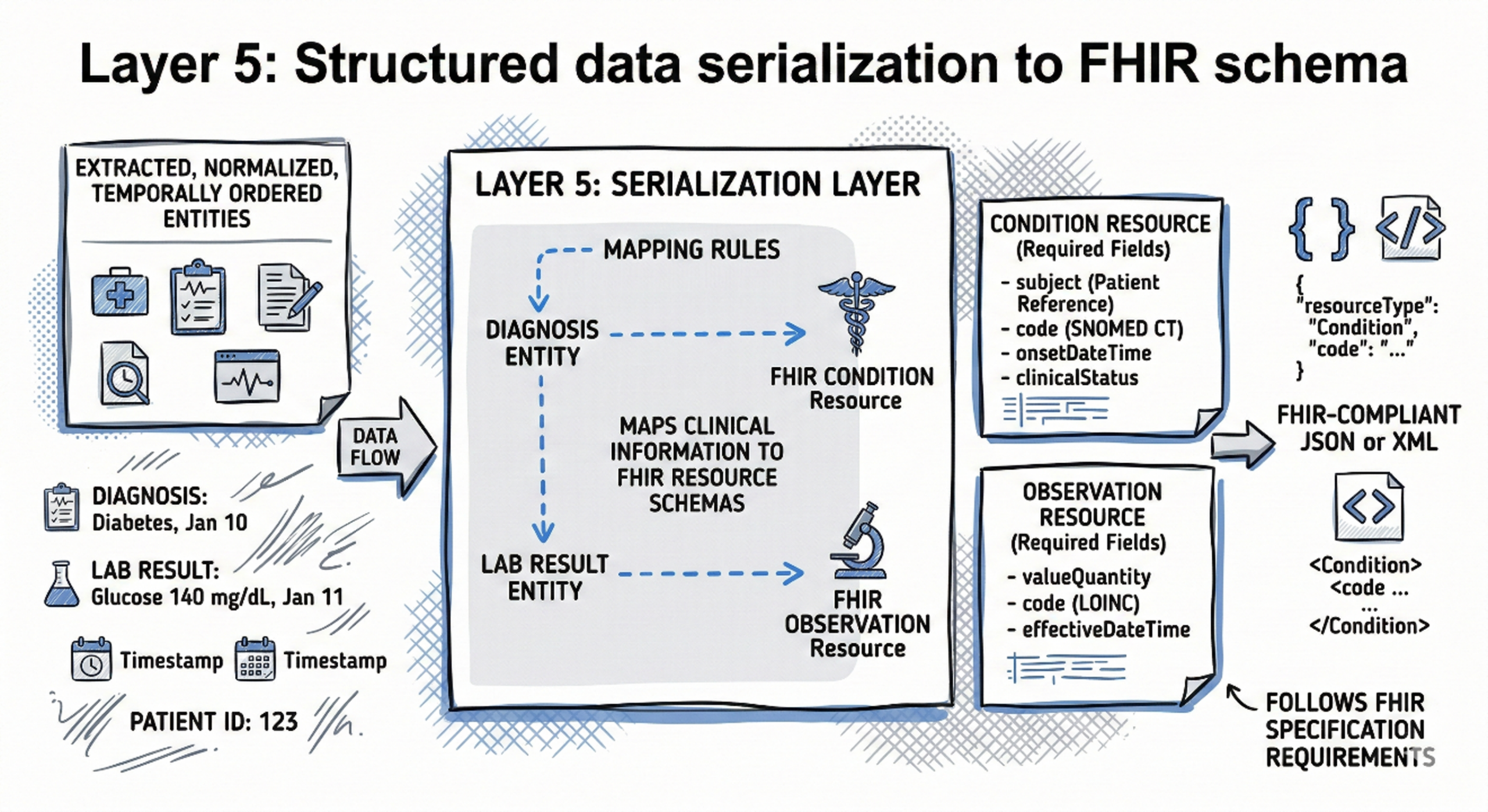
Layer 5: Structured data serialization to FHIR schema
Extracted, normalized, and temporally ordered entities must serialize into FHIR-compliant JSON or XML. A diagnosis becomes a Condition resource with required fields including subject (patient reference), code (SNOMED CT), onsetDateTime, and clinicalStatus. Laboratory results become Observation resources with valueQuantity, code (LOINC), and effectiveDateTime. The serialization layer maps extracted clinical information to FHIR resource schemas following FHIR specification requirements.
Cancer registry teams demonstrate validation requirements for FHIR-ready data. Registries accepting FHIR submissions require data quality meeting regulatory standards. Organizations should implement sampling-based validation comparing automated extraction against expert review, confidence scoring flagging uncertain extractions for human validation, comprehensive audit trails documenting provenance from source notes through FHIR resources, and continuous monitoring detecting accuracy drift requiring model refinement.
Why specialized healthcare NLP enables FHIR-ready extraction
The precision requirements for FHIR resource generation are unforgiving. Incorrect entity extraction, missed contextual signals, or faulty terminology mapping creates FHIR resources misrepresenting patient clinical status, breaking interoperability and creating patient safety risks.
Systematic assessment on 48 medical expert-annotated clinical documents showed Healthcare NLP achieving 96% F1-score for entity detection compared to GPT-4o’s 79%, with GPT-4o completely missing 14.6% of entities versus Healthcare NLP’s 0.9% miss rate.* For FHIR resource generation, a 14.6% miss rate means critical clinical information never reaches interoperable format, quality measures calculate incorrectly, care coordination systems lack essential data, and analytics operate on incomplete patient pictures.
The CLEVER study found medical doctors preferred specialized healthcare natural language processing 45-92% more often than GPT-4o on factuality, clinical relevance, and conciseness.* For FHIR applications where incorrect Condition resources trigger inappropriate clinical decision support or wrong Medication resources cause drug interaction alerts, this factuality gap determines whether systems can be trusted.
Healthcare NLP reduces processing costs by over 80% compared to cloud-based large language model APIs through fixed-cost local deployment.* For organizations converting millions of clinical notes to FHIR resources, per-request API pricing is economically infeasible. On-premise deployment with predictable costs enables sustainable FHIR transformation at enterprise scale.
Production workflow: from clinical note to FHIR resource
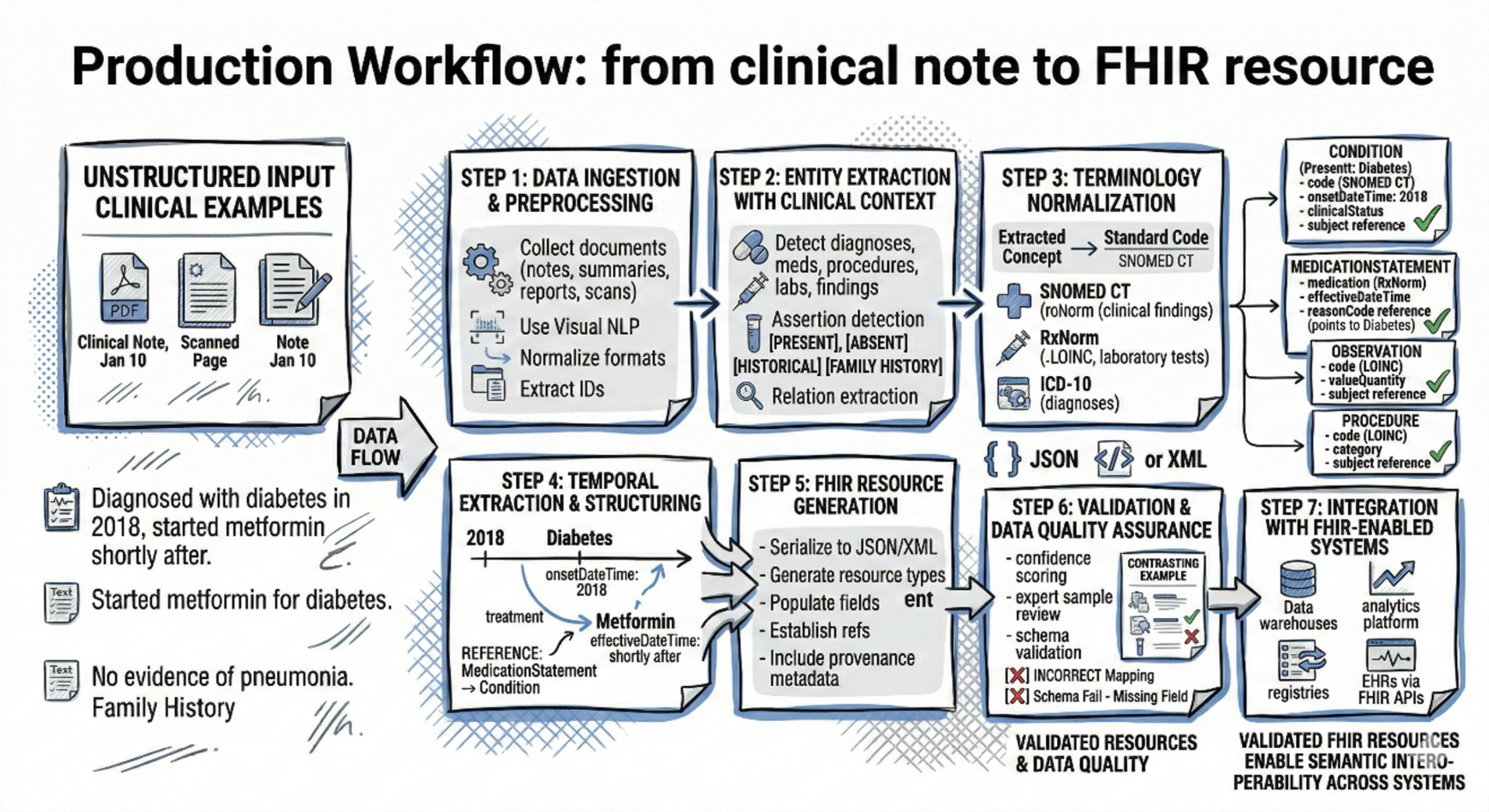
Step 2: Entity extraction with clinical context Healthcare NLP pipelines detect diagnoses, medications, procedures, laboratory tests, anatomical locations, and clinical findings. Assertion detection identifies whether findings are present, absent, historical, or family history. Relation extraction connects medications to conditions, procedures to indications, and observations to clinical context.
Step 3: Terminology normalization Entity resolution maps extracted concepts to standard terminologies: SNOMED CT for clinical findings and conditions, RxNorm for medications, LOINC for laboratory tests and observations, ICD-10 for diagnoses. This standardization enables semantic interoperability across systems.
Step 4: Temporal extraction and structuring Extract dates, temporal relationships, and clinical timelines. “Diagnosed with diabetes in 2018, started metformin shortly after” must structure as: Condition resource with onsetDateTime 2018, MedicationStatement resource with effectiveDateTime shortly after Condition onset, reference linking MedicationStatement to Condition as treatment indication.
Step 5: FHIR resource generation Serialize structured, normalized, temporally ordered data into FHIR-compliant JSON or XML. Generate appropriate resource types: Condition for diagnoses, MedicationStatement for medications, Observation for laboratory results and vital signs, Procedure for interventions, DiagnosticReport for imaging and pathology findings. Populate required fields, establish resource references, include metadata documenting provenance.
Step 6: Validation and data quality assurance Generative AI Lab enables human-in-the-loop validation workflows with audit trails. Organizations should implement confidence scoring, sample-based expert review, FHIR resource validation against schema requirements, and continuous monitoring for extraction accuracy and terminology mapping correctness.
Step 7: Integration with FHIR-enabled systems Load FHIR resources into data warehouses, analytics platforms, registries, or electronic health record systems supporting FHIR APIs. Enable querying, analytics, quality measurement, population health management, and data exchange across organizational boundaries.
Strategic benefits of FHIR-ready AI at scale
Bridging data silos for interoperability: Unstructured notes, legacy reports, and external documents become part of interoperable data fabric. Real Chemistry’s unified view across over 10 datasets demonstrates integration value. Previously siloed information becomes queryable, analyzable, and exchangeable through standardized FHIR resources.
Improved quality measurement and reporting: Richer clinical data extracted from notes supports value-based care initiatives, quality measure calculation, risk adjustment, and population health metrics. Conditions documented only in narrative text become FHIR Condition resources contributing to HEDIS measures and quality reporting.
Enhanced analytics and decision support: Structured, standardized FHIR resources enable predictive modeling, cohort identification, risk stratification, clinical decision support, and real-world evidence generation. Cancer registries demonstrate research value, FHIR-compliant submissions from automated extraction make registry population feasible for resource-constrained organizations.
Scalable data exchange across organizations: FHIR resources facilitate sharing across health systems, registries, research collaborations, and care coordination networks. Standardized format eliminates custom integration development for each data exchange partnership.
Operational efficiency replacing manual abstraction: Automated pipelines reduce labor-intensive chart review, improve data timeliness, and scale across large patient populations. Organizations processing tens of thousands of encounters monthly cannot sustain manual abstraction for FHIR resource generation.
Implementation considerations and common challenges
Clinical language complexity: Ambiguity, abbreviations, context dependencies, and documentation style variation complicate extraction. Generic natural language processing often fails on clinical nuance. Healthcare NLP’s domain-trained models handle medical terminology, negation, temporality, and assertion detection that general-purpose systems miss.
Terminology mapping accuracy: Translating free text to standard codes requires careful normalization and validation. “MI” might mean myocardial infarction, mitral insufficiency, or mental illness depending on clinical context. Entity resolution must disambiguate based on surrounding clinical information.
Document type heterogeneity: Clinical notes, pathology reports, radiology findings, discharge summaries, and scanned historical documents each require specialized handling. Visual NLP processes scanned PDFs and legacy documents. Modular pipelines adapt extraction logic to document types while maintaining consistent FHIR output schemas.
Governance and compliance: Processing protected health information requires HIPAA compliance, audit logging, version control, and secure deployment. De-identification enables privacy-preserving FHIR resource generation when data sharing crosses organizational boundaries.
Ongoing maintenance: Clinical practices, documentation styles, coding standards, and FHIR profiles evolve. Organizations must plan for model updates, terminology mapping maintenance, FHIR schema evolution, and continuous validation. It means ongoing operations, not one-time implementation.
Organizations can explore FHIR-ready AI capabilities through Healthcare NLP for entity extraction and terminology normalization, Visual NLP for scanned document processing, Generative AI Lab for validation workflows, and technical documentation showing implementation patterns. The customer implementations demonstrate production architectures transforming unstructured clinical data into FHIR-compliant resources.
FAQs
What FHIR resources can be generated from unstructured clinical notes?
Healthcare NLP extraction enables generation of multiple FHIR resource types. Diagnoses and clinical findings become Condition resources with SNOMED CT coding. Medications mentioned in notes become MedicationStatement resources with RxNorm codes, dosages, and administration routes. Laboratory test results and vital signs become Observation resources with LOINC codes and quantitative values. Procedures and interventions become Procedure resources. Imaging findings and pathology results become DiagnosticReport resources. Temporal information extracted from narrative text populates onsetDateTime, effectiveDateTime, and other temporal fields. Patient, encounter, and practitioner references link resources establishing clinical context.
How accurate must entity extraction be before FHIR resource generation is reliable?
Systematic benchmarking establishes thresholds: Healthcare NLP achieving 96% F1-score with 0.9% complete miss rate demonstrates production-grade accuracy for FHIR applications. GPT-4o’s 79% F1-score with 14.6% complete miss rate is insufficient as it implies nearly one in seven clinical entities never reaches FHIR resources, creating incomplete patient records and incorrect quality measures. The CLEVER study showing physicians preferring specialized models 45-92% more often reflects the factuality threshold FHIR applications require. Organizations should validate extraction accuracy on institutional data measuring: (1) entity-level precision and recall for conditions, medications, procedures, and observations, (2) assertion detection correctly identifying present versus absent versus historical, (3) relationship extraction linking medications to indications and procedures to findings, (4) terminology mapping to correct SNOMED CT, RxNorm, LOINC, and ICD-10 codes, and (5) temporal extraction accuracy for onset dates and clinical timelines. Cancer registries accepting FHIR submissions demand data quality meeting regulatory standards, requiring comprehensive accuracy validation before automated extraction deploys.
How do organizations handle terminology normalization to multiple standard vocabularies?
Healthcare NLP’s entity resolution supports simultaneous mapping to multiple terminologies. A single diagnosis extraction can map to SNOMED CT for semantic interoperability, ICD-10 for billing and quality measures, and institution-specific coding systems. Medications map to RxNorm for standardization and local formulary codes. Laboratory tests map to LOINC for observation resources and local laboratory information system codes. Organizations should establish mapping priorities based on downstream use cases: FHIR resource exchange across organizations requires SNOMED CT and LOINC, quality measure calculation may require ICD-10, and internal analytics might need local coding preservation. Real Chemistry’s harmonization of over 10 datasets demonstrates multi-terminology requirements: client-specific terminologies must map to standard vocabularies while preserving local codes for internal analytics. Organizations implementing FHIR transformation should plan for ongoing terminology maintenance as SNOMED CT, LOINC, and RxNorm receive quarterly or annual updates requiring mapping validation and potential refinement.
Can FHIR resource generation work with scanned historical documents and PDFs?
Yes, through optical character recognition and specialized document processing. Visual NLP processes scanned pathology reports, radiology documents, historical records, and PDF files that comprise significant portions of healthcare data. The workflow includes: (1) optical character recognition extracting text from scanned images, (2) layout detection identifying document structure and relevant sections, (3) entity extraction from OCR output using Healthcare NLP models, (4) quality validation checking OCR accuracy and extraction confidence, (5) terminology normalization mapping to standard vocabularies, and (6) FHIR resource generation following standard workflow. OCR quality affects extraction accuracy. Organizations should validate on representative scanned documents, implement confidence thresholds for low-quality OCR requiring human review, and prioritize digitally-born documents when available.
How long does it take to implement FHIR-ready AI pipelines?
Implementation timelines depend on scope, data complexity, and existing infrastructure. Organizations can deploy Healthcare NLP pipelines processing standard clinical notes in weeks for initial validation. Comprehensive FHIR transformation requires longer timelines: (1) data source identification and integration planning (4-8 weeks), (2) entity extraction pipeline development using pre-trained models (4-6 weeks), (3) terminology normalization configuration (4-6 weeks), (4) FHIR resource schema mapping and serialization logic (4-8 weeks), (5) validation against institutional gold standard (4-8 weeks), (6) integration with FHIR-enabled systems (4-8 weeks), and (7) production deployment with monitoring (2-4 weeks). Total timeline from planning to production typically ranges from 5-8 months for comprehensive implementations. Organizations should plan phased deployment: start with high-value FHIR resource types like Condition and MedicationStatement for quality measurement, validate accuracy and workflow integration, then expand to additional resource types including Observation, Procedure, and DiagnosticReport. Organizations should plan for continuous enhancement as new document types and use cases emerge.
What is the cost difference between manual abstraction and automated FHIR resource generation?
Manual chart abstraction for FHIR resource generation typically costs $50-150 per patient record depending on complexity, requiring trained abstractors reviewing clinical notes, extracting relevant information, coding to standard terminologies, and populating FHIR resources. For organizations needing FHIR resources for tens of thousands of patients, manual abstraction costs become prohibitive: 100,000 patients at $75 per record costs $7.5M. Automated extraction using Healthcare NLP incurs: (1) Healthcare NLP licensing (annual subscription), (2) infrastructure for processing (cloud or on-premise compute), (3) implementation and validation effort (one-time), and (4) ongoing maintenance (annual). Organizations processing 100,000+ patient records annually typically achieve breakeven within first year, then realize 80-90% cost savings in subsequent years. Healthcare NLP reducing processing costs by over 80% versus cloud APIs through fixed-cost on-premise deployment further improves economics at scale. Additionally, automated extraction enables use cases impossible with manual abstraction: continuous FHIR resource generation as new notes arrive, historical backlog processing, and real-time quality measure updates. These operational capabilities justify investment beyond direct cost comparison.
How do organizations validate FHIR resources generated from unstructured text meet interoperability requirements?
Validation requires multiple layers: (1) FHIR schema validation ensuring resources conform to FHIR specification including required fields, data types, and resource references, (2) terminology binding validation confirming codes use specified value sets so conditions use SNOMED CT, medications use RxNorm, and observations use LOINC, (3) clinical accuracy validation comparing extracted information against source documentation through expert review, (4) semantic interoperability testing verifying resources can be consumed by target systems including electronic health records, registries, and analytics platforms, (5) reference integrity checking ensuring patient, encounter, and practitioner references resolve correctly, and (6) temporal consistency validation confirming onset dates, effective times, and clinical sequencing are logically coherent. Cancer registries reject non-conformant to FIHR resources, requiring comprehensive validation before automated extraction deploys. Organizations should implement Generative AI Lab human-in-the-loop workflows enabling systematic quality assurance, maintain audit trails documenting validation results, and establish continuous monitoring detecting resource quality issues requiring pipeline refinement.
What solutions does John Snow Labs offer for FHIR-ready AI transformation?
John Snow Labs provides comprehensive infrastructure for transforming unstructured clinical data into FHIR-compliant resources. Healthcare NLP includes over 2,800 pre-trained clinical models for entity extraction, assertion detection, relationship extraction, and entity resolution mapping to SNOMED CT, ICD-10, LOINC, and RxNorm, the standard terminologies FHIR resources require. Visual NLP processes scanned documents and PDF reports through optical character recognition and layout detection. Medical LLM provides specialized clinical reasoning for complex extraction tasks. Generative AI Lab enables human-in-the-loop validation workflows with audit trails ensuring FHIR resource quality. De-identification provides HIPAA-compliant processing when FHIR resources are exchanged across organizational boundaries. These integrate with Databricks, AWS, Azure, and on-premise environments supporting flexible deployment. Organizations can explore live demonstrations showing entity extraction and terminology normalization, review customer implementations including Real Chemistry and cancer registry teams, or access technical documentation for FHIR transformation architecture patterns and implementation guidance.




























