Boosting Efficiency and Accuracy in Healthcare NLP Tasks Using Healthcare-Specific Fine-Tuned LLMs and New Medical Assertion Detection Frameworks
Highlights
- Introducing a brand new
LLMLoaderannotator to load and run large language models in gguf We also announce 9 LLMs at various sizes and quantizations (3x small size medical summarizer and QA model, 3x medium size general model, and 3x small size zero shot entity extractor) - Introducing a brand new
FewshotAssertionClassifierannotator to train assertion detection models using a few samples with better accuracy - Introducing a rule-based
ContextualAssertionannotator to detect assertion status using patterns and rules without any training or annotation - Introducing
VectorDBPostProcessorannotator to filter and sort the document splits returned by vector databases in an RAG application - Introducing
ContextSplitAssemblerannotator to assemble the document post-processed splits as a context into an LLM stage in an RAG application - SNOMED entity resolver model for veterinarythe domain
- Voice of the Patients (VOP) named entity recognition (NER) model
- New rule-based entity matcher models to customize De-Identification pipelines
- New NER, assertion, relation extraction, and classification models to identify
AlcoholandSmokingrelated Medical Entities - New NER and assertion models to extract
Menopauserelated entities - Clinical document analysis with one-liner pretrained pipelines for specific clinical tasks and concepts
- The formal release of oncological assertion status detection and relation extraction models
- 11 new fine-tuned sentence embedding models finetuned with medical assertion datasets
- Significantly faster vector-db-based entity resolution models than existing Sentence Entity Resolver models
- RxNorm code mapping benchmarks and cost comparisons: Healthcare NLP, GPT-4, and Amazon
- New blog posts on using NLP in opioid research and healthcare: harnessing NLP, knowledge graphs, and regex techniques for critical insights
- New notebooks for medication and resolution concept
- Updated Udemy MOOC (our online courses) notebooks
- Various core improvements; bug fixes, enhanced overall robustness and reliability of Spark NLP for Healthcare
- Updated notebooks and demonstrations for making Spark NLP for Healthcare easier to navigate and understand
- New JSL vs GPT-4 Demo to showcase accuracy differences between Healthcare NLP and GPT-4 for information extraction tasks
- The addition and update of numerous new clinical models and pipelines continue to reinforce our offering in the healthcare domain
If you would like to check the whole release notes, you can see it here.
Introducing a Brand New LLMLoader Annotator to Load and Run Large Language Models in GGUF format
LLMLoader is designed to interact with a LLMs that are converted into gguf format. This module allows using John Snow Labs’ licensed LLMs at various sizes that are finetuned on medical context for certain tasks. It provides various methods for setting parameters, loading models, generating text, and retrieving metadata. The LLMLoader includes methods for setting various parameters such as input prefix, suffix, cache prompt, number of tokens to predict, sampling techniques, temperature, penalties, and more. Overall, the LLMLoader provides a flexible and extensible framework for interacting with language models in a Python and Scala environment using PySpark and Java.
Example:
from sparknlp_jsl.llm import LLMLoader
llm_loader_pretrained = LLMLoader(spark).pretrained("JSL_MedS_q16_v1", "en", "clinical/models")
llm_loader_pretrained.generate("What is the indication for the drug Methadone?")
Result:
Methadone is used to treat opioid addiction. It is a long-acting opioid agonist that is used to help individuals who are addicted to short-acting opioids such as heroin or other illicit opioids. It is also used to treat chronic pain in patients who have developed tolerance to other opioids.
Introducing a Brand New FewshotAssertionClassifier Annotator to Train Assertion Detection Models Using a Few Samples with Better Accuracy
The newly refactored FewShotAssertionClassifierModel and FewShotAssertionClassifierApproach simplify assertion annotation in clinical and biomedical texts. These models deliver precise assertion annotations by leveraging sentence embeddings and integrate seamlessly with any SparkNLP sentence embedding model.
A key feature is the FewShotAssertionSentenceConverter, an annotator that formats documents/sentences and NER chunks for assertion classification, requiring an additional step in the pipeline.
This comprehensive approach significantly enhances the extraction, analysis, and processing of assertion-related data, making it an indispensable tool for healthcare text annotation.
The following table demonstrates the enhanced results achieved using the FewShot Assertion model compared to the traditional AssertionDL model across various datasets. The FewShot Assertion model showcases significant accuracy improvement scores, particularly in complex medical domains.
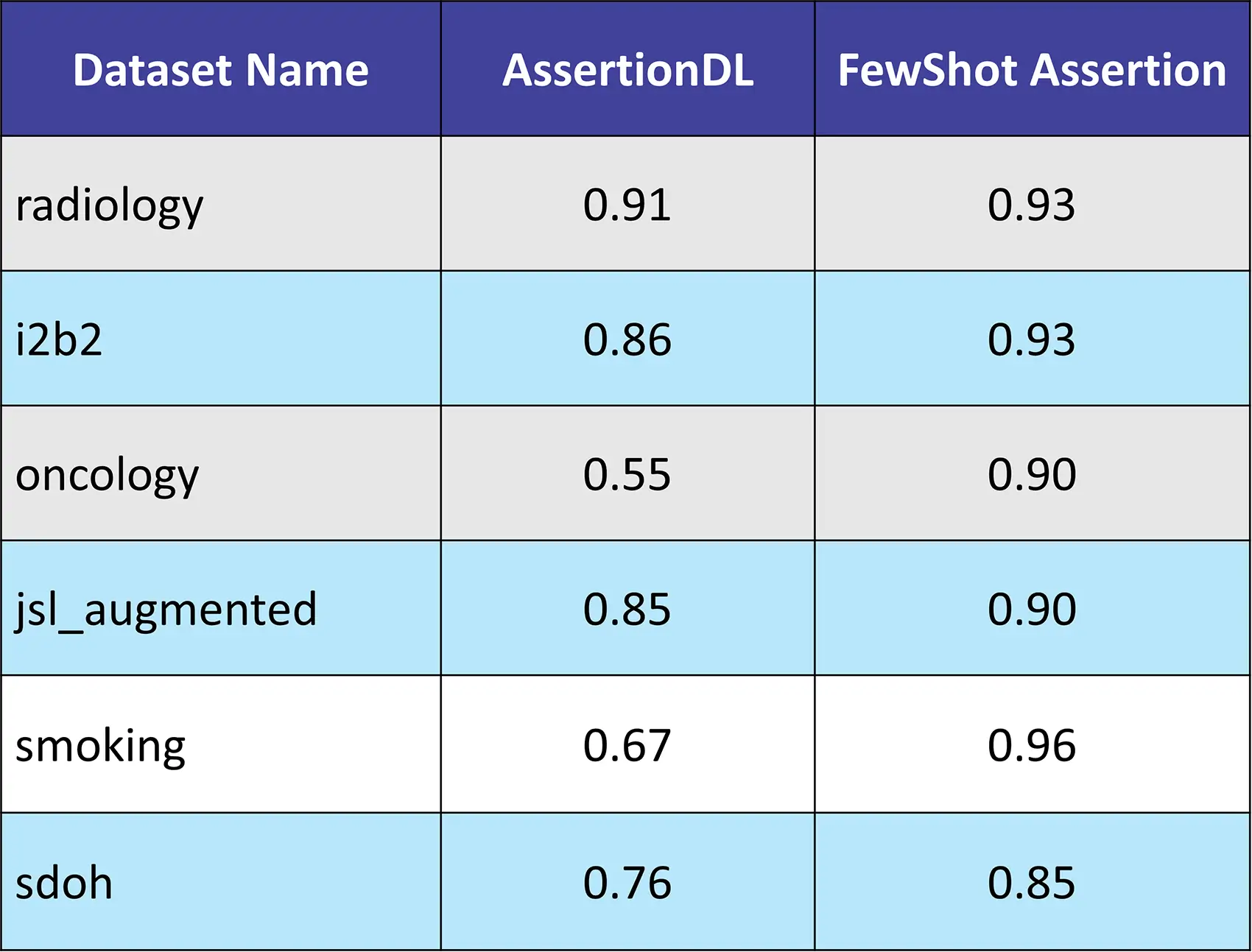
Accuracy Table
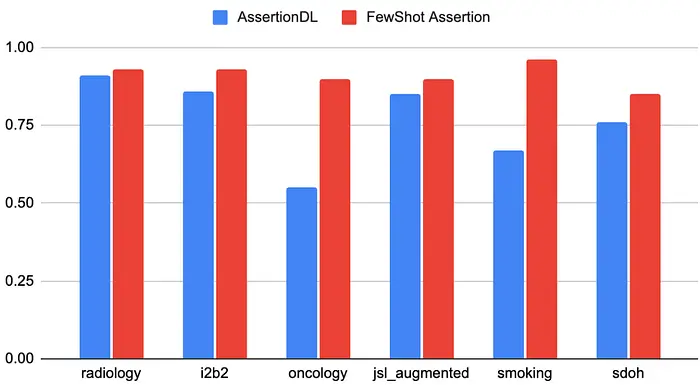
Accuracy Comparison Table
FewShot Assertion Models:
- fewhot_assertion_jsl_e5_base_v2_jsl:
Present,Absent,Possible,Planned,Past,Family,Hypothetical,SomeoneElse - fewhot_assertion_i2b2_e5_base_v2_i2b2:
absent,associated_with_someone_else,conditional,hypothetical,possible,present - fewhot_assertion_sdoh_e5_base_v2_sdoh:
Absent,Past,Present,Someone_Else,Hypothetical,Possible - fewhot_assertion_smoking_e5_base_v2_smoking:
Present,Absent,Past - fewhot_assertion_oncology_e5_base_v2_oncology:
Absent,Past,Present,Family,Hypothetical,Possible - fewhot_assertion_radiology_e5_base_v2_radiology:
Confirmed,Negative,Suspected
Example:
few_shot_assertion_converter = FewShotAssertionSentenceConverter()\
.setInputCols(["sentence","token", "ner_jsl_chunk"])\
.setOutputCol("assertion_sentence")
e5_embeddings = E5Embeddings.pretrained("e5_base_v2_embeddings_medical_assertion_oncology", "en", "clinical/models")\
.setInputCols(["assertion_sentence"])\
.setOutputCol("assertion_embedding")
few_shot_assertion_classifier = FewShotAssertionClassifierModel()\
.pretrained("fewhot_assertion_oncology_e5_base_v2_oncology", "en", "clinical/models")\
.setInputCols(["assertion_embedding"])\
.setOutputCol("assertion")
sample_text= """The patient is suspected to have colorectal cancer. Her family history is positive for other cancers. The result of the biopsy was positive. A CT scan was ordered to rule out metastases."""
Please check the FewShot Assertion Classifier Notebook for more information.
Introducing a Rule-Based ContextualAssertion Annotator to Detect Assertion Status Using Patterns and Rules without any Training or Annotation
Introducing Contextual Assertion which identifies contextual cues within text data, such as negation, uncertainty, etc. It is used for clinical assertion detection, etc. It annotates text chunks with assertions based on configurable rules, prefix and suffix patterns, and exception patterns.
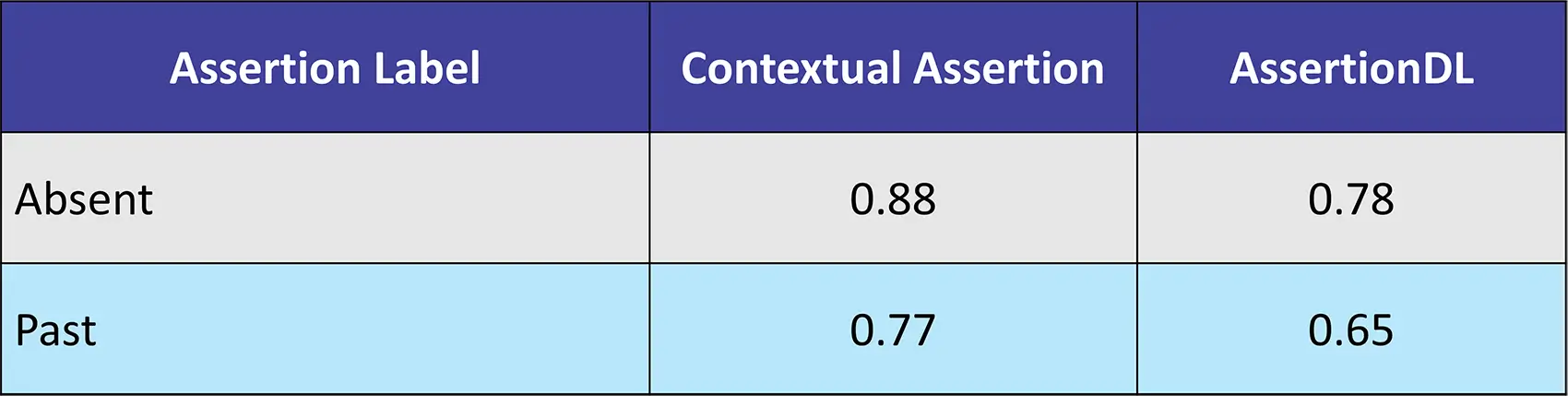
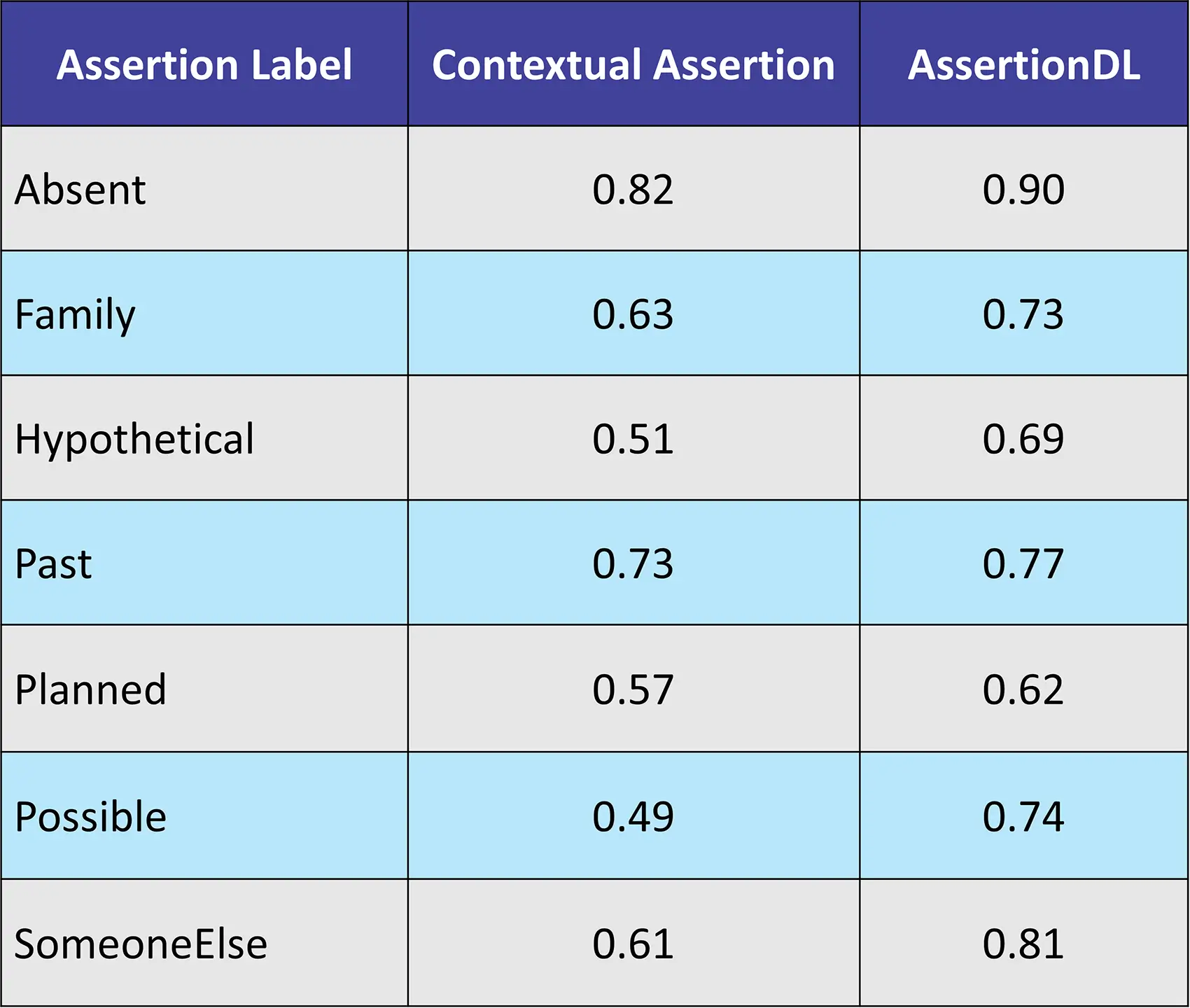
Dataset: 253 Clinical Texts from in-house dataset
Contextual Assertion, a powerful component within Healthcare NLP, extends beyond mere negation detection. Its ability to identify and classify various contextual cues, including uncertainty, temporality, and sentiment, empowers healthcare professionals to extract deeper meaning from complex medical records.
Contextual Assertion Models:
- contextual_assertion_someone_else: Identifies contextual cues within text data to detect
someone elseassertions - contextual_assertion_absent: Identifies contextual cues within text data to detect
absentassertions - contextual_assertion_past: Identifies contextual cues within text data to detect
pastassertions
Example:
contextual_assertion = ContextualAssertion() \
.setInputCols(["sentence", "token", "ner_chunk"]) \
.setOutputCol("assertion") \
.setPrefixKeywords(["no", "not"]) \
.setSuffixKeywords(["unlikely", "negative", "no"]) \
.setPrefixRegexPatterns(["\\b(no|without|denies|never|none|free of|not include)\\b"]) \
.setSuffixRegexPatterns(["\\b(free of|negative for|absence of|not|rule out)\\b"]) \
.setExceptionKeywords(["without"]) \
.setExceptionRegexPatterns(["\\b(not clearly)\\b"]) \
.addPrefixKeywords(["negative for", "negative"]) \
.addSuffixKeywords(["absent", "neither"]) \
.setCaseSensitive(False) \
.setPrefixAndSuffixMatch(False) \
.setAssertion("absent") \
.setScopeWindow([2, 2])\
.setIncludeChunkToScope(True)
sample_text = """Patient resting in bed. Patient given azithromycin without any difficulty. Patient has audible wheezing, states chest tightness.
No evidence of hypertension. Patient denies nausea at this time. zofran declined. Patient is also having intermittent sweating associated with pneumonia.
"""
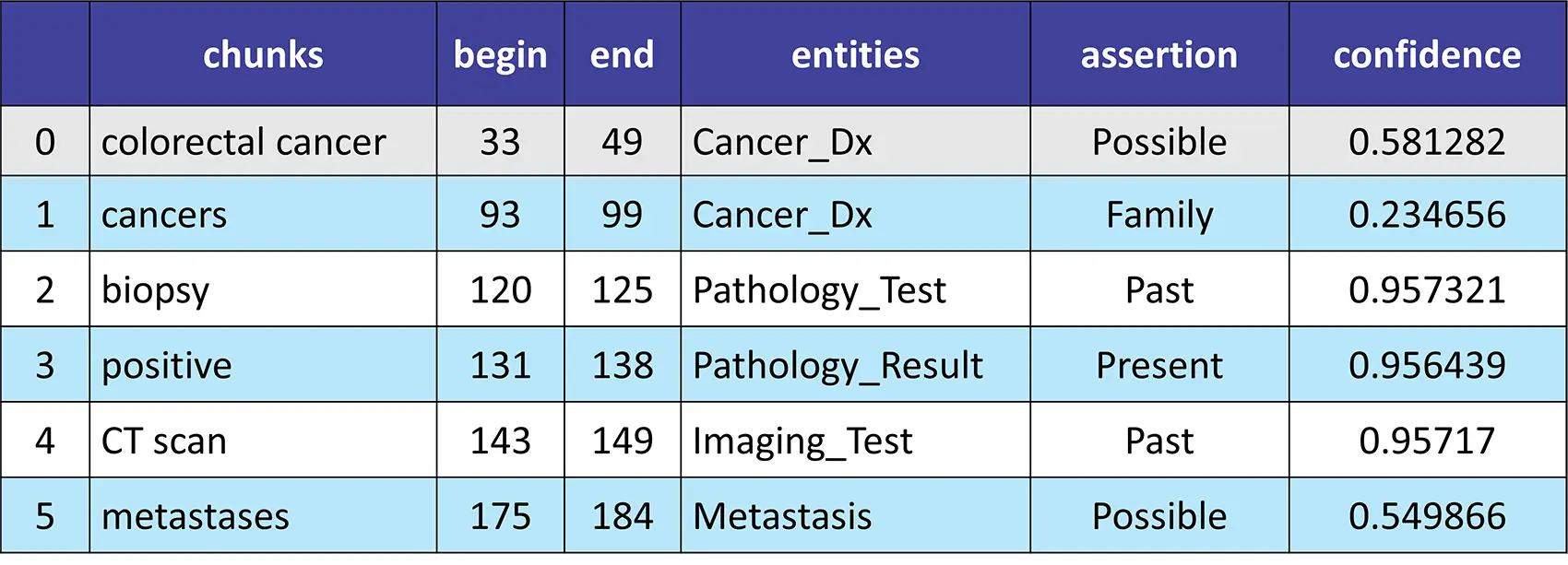
Result:
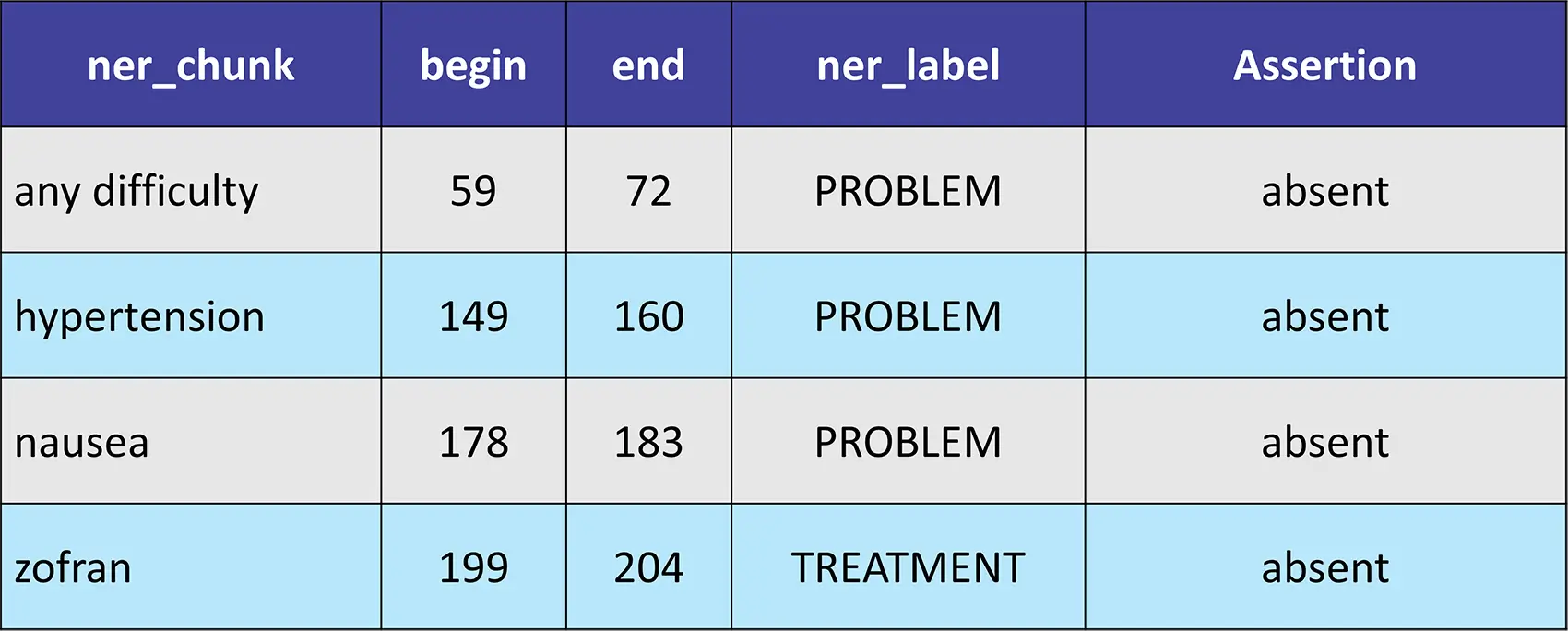
Introducing VectorDBPostProcessor Annotator to Filter and Sort the Document Splits Returned by VectorDB in a RAG Application
The VectorDBPostProcessor is a powerful tool designed to filter and sort output from the VectorDBModel (our own VectorDB implementations will be released soon). This processor refines VECTOR_SIMILARITY_RANKINGS input annotations and outputs enhanced VECTOR_SIMILARITY_RANKINGS annotations based on specified criteria.
Key Parameters:
filterBy(str): Select and prioritize filter options (metadata, diversity_by_threshold). Options can be given as a comma-separated string, determining the filtering order. Default: metadatasortBy(str): Select sorting option (ascending, descending, lost_in_the_middle, diversity). Default: ascendingcaseSensitive(bool): Determines if string operators’ criteria are case-sensitive. Default: FalsediversityThreshold(float): Sets the threshold for the diversity_by_threshold filter. Default: 0.01maxTopKAfterFiltering(int): Limits the number of annotations returned after filtering. Default: 20allowZeroContentAfterFiltering(bool): Determines whether zero annotations are allowed after filtering. Default: False
This processor ensures precise and customizable annotation management, making it an essential component for advanced data processing workflows.
Example
post_processor = VectorDBPostProcessor() \
.setInputCols("vector_db") \
.setOutputCol("post") \
.setSortBy("ascending")
.setMaxTopKAfterFiltering(5)
.setFilterBy("metadata") \
.setMetadataCriteria([
{"field": "pubdate", "fieldType": "date", "operator": "greater_than", "value": "2017 May 11", "dateFormats": ["yyyy MMM dd", "yyyy MMM d"], "converterFallback": "filter"},
{"field": "distance", "fieldType": "float", "operator": "less_than", "value": "0.5470"},
{"field": "title", "fieldType": "string", "operator": "contains", "matchMode": "any", "matchValues": ["diabetes", "immune system"]}
])
Introducing ContextSplitAssembler Annotator to Assemble the Document Post-processed Splits as a Context into an LLM Stage in a RAG Application
The ContextSplitAssembler is a versatile tool designed to work seamlessly with vector databases (our own VectorDB implementations will be released soon) and VectorDBPostProcessor. It combines and organizes annotation results with customizable delimiters and optional splitting.
Key Parameters:
joinString(str): Specifies the delimiter string inserted between annotations when combining them into a single result. Ensures proper separation and organization. Default: “ “explodeSplits(bool): Determines whether to split the annotations into separate entries. Default: False
This assembler enhances the management and presentation of annotations, making it an essential tool for advanced data processing workflows.
Example:
context_split_assembler = ( ContextSplitAssembler()
.setInputCols("vector_db")
.setOutputCol("document")
.setJoinString("\n")
.setExplodeSplits(False))
Please check the VectorDB and PostProcessor for RAG Generative AI Notebook for more information.
SNOMED Entity Resolver Model for Veterinary Domains
This advanced model facilitates the mapping of veterinary-related entities and concepts to SNOMED codes using sbiobert_base_cased_mli Sentence BERT embeddings. It is trained with an enhanced dataset derived from the sbiobertresolve_snomed_veterinary_wip model. The model ensures precise and reliable resolution of veterinary terms to standardized SNOMED codes, aiding in consistent and comprehensive veterinary data documentation and analysis.
Example:
snomed_resolver = SentenceEntityResolverModel.pretrained("sbiobertresolve_snomed_veterinary", "en", "clinical/models") \
.setInputCols(["sentence_embeddings"]) \
.setOutputCol("snomed_code")\
.setDistanceFunction("EUCLIDEAN")
text = "The veterinary team is closely monitoring the patient for signs of lymphoblastic lymphoma, a malignant neoplasm of lymphoid origin. They are also treating the patient's osteoarthritis, a degenerative joint disease. Additionally, the team is vigilantly observing the facility for potential outbreaks of mink distemper."
Result:
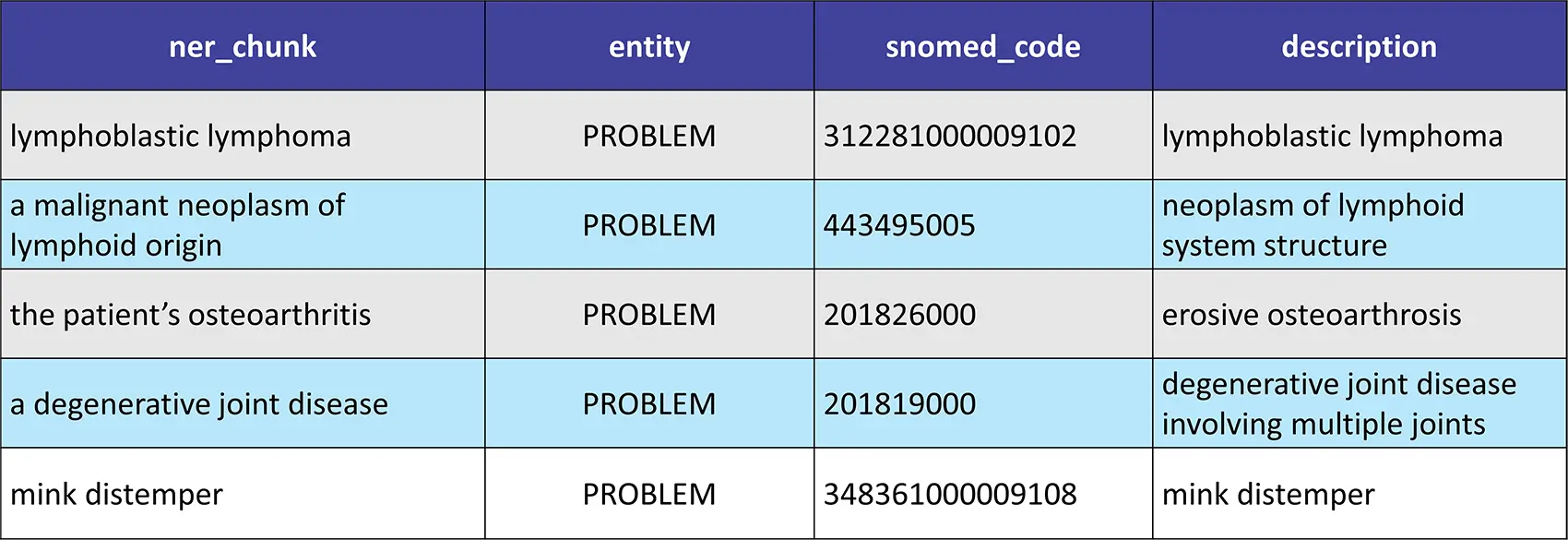
Voice of the Patients Named Entity Recognition (NER) Model
The Voice of the Patients NER Model is designed to extract healthcare-related terms from patient-generated documents. This model processes the natural language used by patients to identify and categorize medical terms, facilitating better understanding and documentation of patient-reported information.
Example
ner_model = MedicalNerModel.pretrained("ner_vop_v2", "en", "clinical/models")\
.setInputCols(["sentence", "token", "embeddings"])\
.setOutputCol("ner")
sample_text = "Hello,I'm 20 year old girl. I'm diagnosed with hyperthyroid 1 month ago. I was feeling weak, light headed,poor digestion, panic attacks, depression, left chest pain, increased heart rate, rapidly weight loss, from 4 months."
Result:
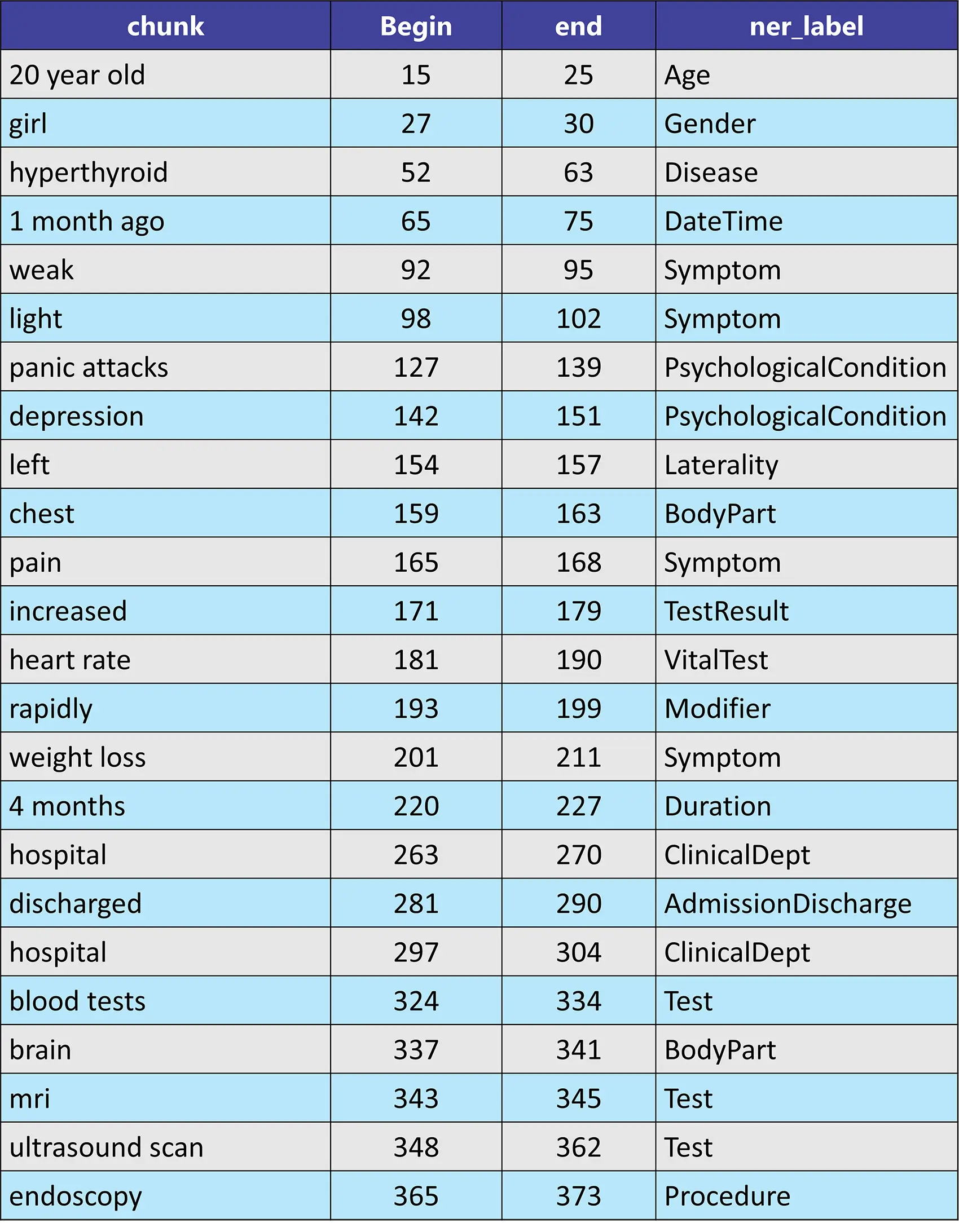
Please check the model card for more information.
Major Update on Healthcare NLP: New Rule-Based Entity Matcher Models to Customise De-IDentification Pipelines
We introduce a suite of text and regex matchers, specifically designed to enhance the deidentification and clinical document understanding process with rule-based methods.
- cancer_diagnosis_matcher: This model extracts cancer diagnoses in clinical notes using a rule-based
TextMatcherInternal - country_matcher: This model extracts countries in clinical notes using a rule-based
TextMatcherInternal - email_matcher: This model extracts emails in clinical notes using a rule-based
RegexMatcherInternal - phone_matcher: This model extracts phone entities in clinical notes using a rule-based
RegexMatcherInternal - state_matcher: This model extracts states in clinical notes using a rule-based
RegexMatcherInternal - zip_matcher: This model extracts zip codes in clinical notes using a rule-based RegexMatcherInternal
- city_matcher: This model extracts city names in clinical notes using a rule-based
TextMatcherInternal
Example:
text_matcher = TextMatcherInternalModel.pretrained("cancer_diagnosis_matcher", "en", "clinical/models") \
.setInputCols(["sentence", "token"])\
.setOutputCol("cancer_dx")\
.setMergeOverlapping(True)
sample_text = """A 65-year-old woman had a history of debulking surgery, bilateral oophorectomy with omentectomy, total anterior hysterectomy with radical pelvic lymph nodes dissection due to ovarian carcinoma (mucinous-type carcinoma, stage Ic) 1 year ago. The patient's medical compliance was poor and failed to complete her chemotherapy (cyclophosphamide 750 mg/m2, carboplatin 300 mg/m2). Recently, she noted a palpable right breast mass, 15 cm in size which nearly occupied the whole right breast in 2 months. Core needle biopsy revealed metaplastic carcinoma. Neoadjuvant chemotherapy with the regimens of Taxotere (75 mg/m2), Epirubicin (75 mg/m2), and Cyclophosphamide (500 mg/m2) was given for 6 cycles with poor response, followed by a modified radical mastectomy (MRM) with dissection of axillary lymph nodes and skin grafting. Postoperatively, radiotherapy was done with 5000 cGy in 25 fractions. The histopathologic examination revealed a metaplastic carcinoma with squamous differentiation associated with adenomyoepithelioma."""
Result:
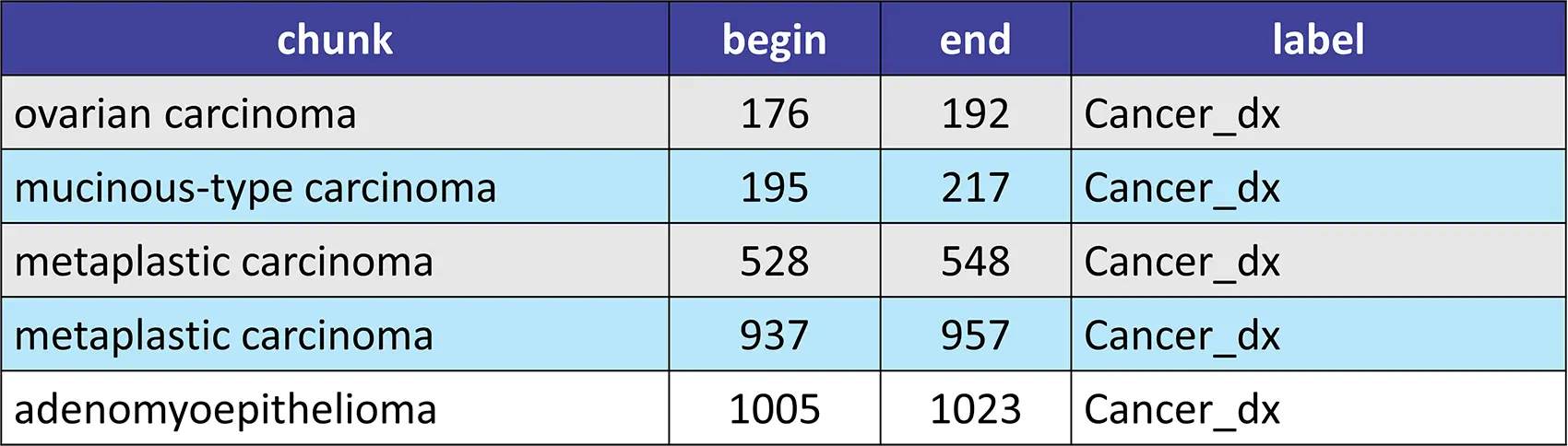
New NER, Assertion, Relation Extraction, and Classification Models to Identify Alcohol and Smoking Related Medical Entities
A suite of models designed for the identification and analysis of alcohol and smoking related entities in text data. These models include Named Entity Recognition (NER), assertion status, relation extraction, and classification, providing a comprehensive toolkit for analyzing substance use information.
NER Model:

ner_alcohol_smoking
Assertion Models:
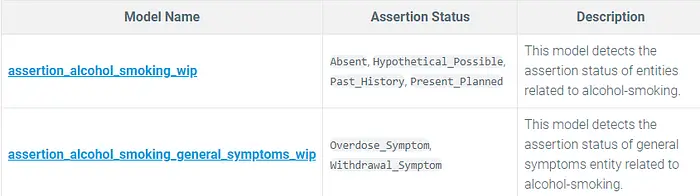
assertion_alcohol_smoking_wip and assertion_alcohol_smoking_general_symptoms_wip
Relation Extraction Model:

re_alcohol_smoking_clinical_wip
Classification Models:
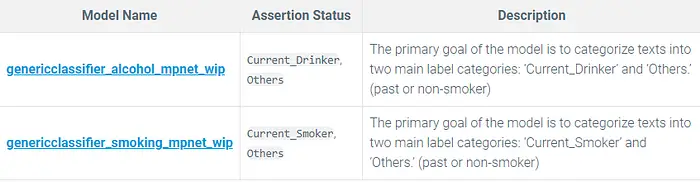
genericclassifier_alcohol_mpnet_wip and genericclassifier_smoking_mpnet_wip
New NER and Assertion Models to Extract Menopause Related Entities
A set of sophisticated models aimed at extracting and analyzing menopause-related entities in text data. These models include a Named Entity Recognition (NER) model and assertion models, which identify and determine the status of various menopause-related terms, aiding in comprehensive menopause data analysis.
NER Model:
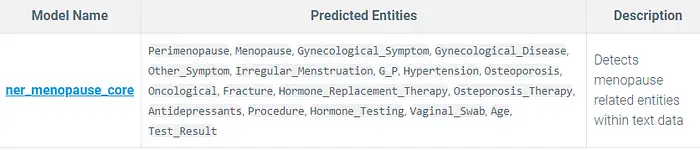
ner_menopause_core
Assertion Model:
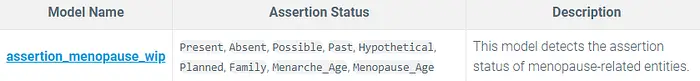
assertion_menopause_wip
Clinical Document Analysis with One-Liner Pretrained Pipelines for Specific Clinical Tasks and Concepts
We introduce a suite of advanced, hybrid pretrained pipelines, specifically designed to streamline the clinical document analysis process. These pipelines are built upon multiple state-of-the-art (SOTA) pretrained models, delivering a comprehensive solution for quickly extracting vital information.
- ner_deid_context_nameAugmented_pipeline: In this pipeline, there are
ner_deid_generic_augmented,ner_deid_subentity_augmented,ner_deid_name_multilingual_clinicalNER models and several <ContextualParser,RegexMatcher, andTextMatcher</code models were used. - ner_profiling_vop: This pipeline can be used to simultaneously evaluate various pre-trained named entity recognition (NER) models, enabling comprehensive analysis of text data pertaining to patient perspectives and experiences, also known as the “Voice of Patients”.
- ner_profiling_sdoh:This pipeline can be used to simultaneously evaluate various pre-trained named entity recognition (NER) models, enabling comprehensive analysis of text data pertaining to the social determinants of health (SDOH). When you run this pipeline over your text, you will end up with the predictions coming out of each pretrained clinical NER model trained with the
embeddings_clinical, which are specifically designed for clinical and biomedical text.
Please check the Task Based Clinical Pretrained Pipelines Notebook for more information.
Formal Release of Oncological Assertion Status Detection and Relation Extraction Models
We are releasing the formal version of the “work-in-progress (WIP)” assertion status detection and relation extraction models in the Oncology domain.
You can check the formal versions of these models from here.
11 New Fine-Tuned Sentence Embedding Models finetuned with medical assertion datasets
Discover our new fine-tuned transformer-based sentence embedding models, meticulously trained on a curated list of clinical and biomedical datasets. These models are specifically optimized for Few-Shot Assertion tasks but are versatile enough to be utilized for other applications, such as Classification and Retrieval-Augmented Generation (RAG). Our collection offers precise and reliable embeddings tailored for various medical domains, significantly enhancing the extraction, analysis, and processing of assertion-related data in healthcare texts.
You can check the models and a sample of their usage from here.
Significantly Faster Vector-DB Based Entity Resolution Models Than Existing Sentence Entity Resolver Models
We have developed vector database-based entity resolution models that are 10x faster on GPU and 2x as fast on CPU compared to the existing Sentence Entity Resolver models.
NOTE: These models are not available on the Models Hub page yet and cannot be used like the other Spark NLP for Healthcare models. They will be integrated into the marketplace and made available there soon.
RxNorm Code Mapping Benchmarks and Cost Comparisons: Healthcare NLP, GPT-4, and Amazon Comprehend Medical
We have prepared an accuracy benchmark and the cost analysis between Healthcare NLP, GPT-4, and Amazon Comprehend Medical for mapping medications to their RxNorm terms. Here are the notes:
- For the ground truth dataset, we used 79 in-house clinical notes annotated by the medical experts of John Snow Labs.
- Healthcare NLP: We used
sbiobertresolve_rxnorm_augmentedandbiolordresolve_rxnorm_augmentedmodels for this benchmark. These models can return up to 25 closest results sorted by their distances. - GPT-4: Both GPT-4 (Turbo) and GPT-4o models are used. According to the official announcement, the performance of GPT-4 and GPT-4o is almost identical, and we used both versions for the accuracy calculation. Additionally, the GPT-4 returns only one result, which means you will see the same results in both evaluation approaches.
- Amazon Comprehend Medical: The RxNorm tool of this service is used, and it returns up to 5 closest matches sorted by their distances.
- We adopted two approaches for evaluating these tools, given that the model outputs may not precisely match the annotations:
- Top-3:Compare the annotations to see if they appear in the first three results.
- Top-5:Compare the annotations to see if they appear in the first five results.
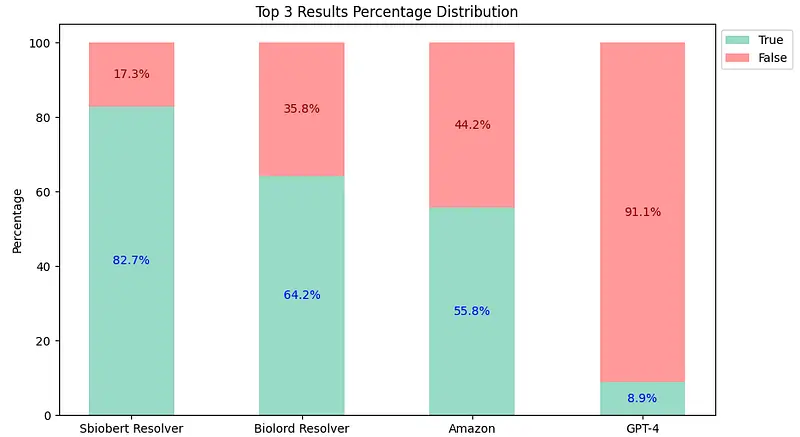
Top-3 Comparison Results
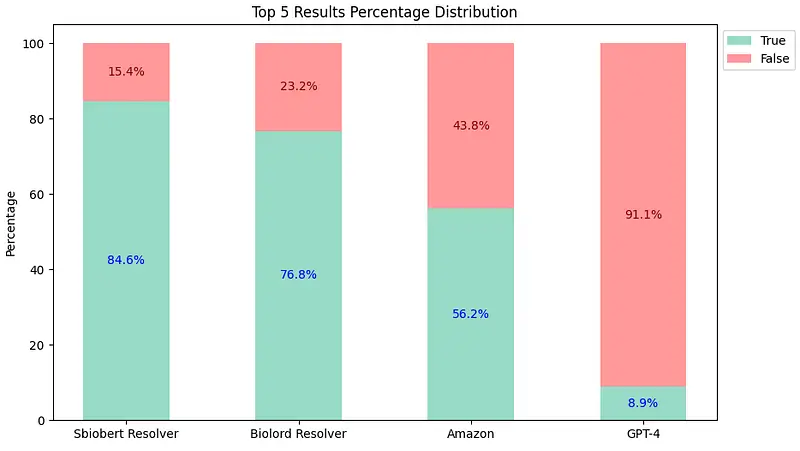
Top-5 Comparison Results
Conclusion:
Based on the evaluation results:
- The
sbiobertresolve_rxnorm_augmentedmodel of Healthcare NLP consistently provides the most accurate results in each top_k comparison. - The
biolordresolve_rxnorm_augmentedmodel of Healthcare NLP outperforms Amazon Comprehend Medical and GPT-4 in mapping terms to their RxNorm codes. - The GPT-4could only return one result, reflected similarly in both charts and has proven to be the least accurate.
If you want to process 1M documents and extract RxNorm codes for medication entities (excluding the NER stage), the total cost:
- With Healthcare NLP is about $4,500, including the infrastructure costs.
- $24,250 with Amazon Comprehend Medical
- $44,000 with the GPT-4 (Turbo) and $22,000 with the GPT-4o
Therefore, Healthcare NLP is almost 5 times cheaper than its closest alternative, not to mention the accuracy differences (Top 3: Healthcare NLP 82.7% vs Amazon 55.8% vs GPT-4 8.9%).
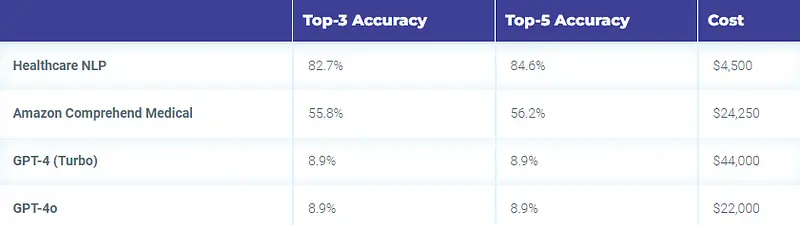
Accuracy & Cost Table
If you want to see more details, please check Benchmarks Page and State-of-the-art RxNorm Code Mapping with NLP: Comparative Analysis between the tools by John Snow Labs, Amazon, and GPT-4 blog post.
New Blogposts on Using NLP in Opioid Research and Healthcare: Harnessing NLP, Knowledge Graphs, and Regex Techniques for Critical Insights
Explore the latest developments in healthcare NLP and Knowledge Graphs through our new blog posts, where we take a deep dive into the innovative technologies and methodologies transforming the medical field. These posts offer insights into how the latest tools are being used to analyze large amounts of unstructured data, identify critical medical assets, and extract meaningful patterns and correlations. Learn how these advances are not only improving our understanding of complex health issues but also contributing to more effective prevention, diagnosis, and treatment strategies.
- Harnessing the Power of NLP and Knowledge Graphs for Opioid Research discusses how Natural Language Processing (NLP) and Knowledge Graphs (KG) are transforming opioid research. By using NLP to process large volumes of unstructured medical data and employing Knowledge Graphs to map intricate relationships, researchers can achieve greater insights into the opioid crisis.
- Extracting Critical Insights on Opioid Use Disorder with Healthcare NLP Models discusses how John Snow Labs’ Healthcare NLP models are transforming the extraction of crucial insights on opioid use disorder. These advanced NLP techniques efficiently identify and categorize medical terminology related to opioid addiction, improving clinical understanding and treatment strategies.
- Extract Medical Named Entities with Regex in Healthcare NLP at Scale explains that the RegexMatcherInternal class employs regular expressions to detect and associate specific text patterns with predefined entities like dates, SSNs, and email addresses. This method facilitates targeted entity extraction by matching text patterns to these predefined entities.
- Extracting Medical Named Entities with Healthcare NLP’s EntityRulerInternal explains that EntityRulerInternal in Spark NLP extracts medical entities from text using regex patterns or exact matches defined in JSON or CSV files. This post explains how to set it up and use it in a Healthcare NLP pipeline, with practical examples.
- Using Contextual Assertion for Clinical Text Analysis: A Comprehensive Guide dive into leveraging Healthcare NLP, a robust NLP library, for clinical text analysis, emphasizing the role of Contextual Assertion. Contextual Assertion markedly enhances the accuracy of detecting negation, possibility, and temporality in medical records. It surpasses deep learning-based assertion status detection in accurately categorizing health conditions. Benchmark comparisons reveal an average F1 score improvement of 10–15%, highlighting the superior precision and reliability of Contextual Assertion in healthcare data analysis.
- State-of-the-art RxNorm Code Mapping with NLP: Comparative Analysis between the tools by John Snow Labs, Amazon, and GPT-4 compares RxNorm code mapping accuracy and a price analysis between John Snow Labs, GPT-4, and Amazon.
New Notebooks for Medication and Resolutions Concept
To better understand the Medication and Resolutions Concept, the following notebooks have been developed:
- New Clinical Medication Use Case notebook: This notebook is designed to extract and analyze medication information from a clinical dataset. Its purpose is to identify commonly used medications, gather details on dosage, frequency, strength, and route, determine current and past usage, understand pharmacological actions, identify treatment purposes, retrieve relevant codes (RxNorm, NDC, UMLS, SNOMED), and find associated adverse events.
- New Resolving Medical Terms to Terminology Codes Directly notebook: In this notebook, you will find how to optimize the process to get SentenceEntityResolverModel model outputs.
- New Analyse Veterinary Documents with Healthcare NLP notebook: In this notebook, we use Spark NLP for Healthcare to process veterinary documents. We focus on Named Entity Recognition (NER) to identify entities, Assertion Status to confirm their condition, Relation Extraction to understand their relationships, and Entity Resolution to standardize terms. This helps us efficiently extract and analyze critical information from unstructured veterinary texts.
Updated Udemy MOOC (Our Online Courses) Notebooks
Recently updated Udemy MOOC (Massive Online Course) notebooks that focus on using Spark NLP annotators for healthcare applications. These notebooks provide practical examples and exercises for learning how to implement and utilize various Spark NLP tools and techniques specifically designed for processing and analyzing healthcare-related text data. The update might include new features, improvements, or additional content to enhance the learning experience for students and professionals in the healthcare field.
Please check the Spark_NLP_Udemy_MOOC folder for the all Healthcare MOOC Notebooks.
Various Core Improvements; Bug Fixes, Enhanced Overall Robustness, and Reliability of Spark NLP for Healthcare
- Resolved broken links in healthcare demos
- Added a unique ID field for each entity into the result of the
pipeline_ouput_parsermodule - Fixed deidentification AGE obfuscation hanging issue
- Added DatasetInfo parameter into the
MedicalNERModelannotator
Updated Notebooks And Demonstrations For making Spark NLP For Healthcare Easier To Navigate And Understand
- New Clinical Medication Use Case notebook
- New Resolving Medical Terms to Terminology Codes Directly notebook
- New Contextual Assertionnotebook
- New VectorDB and PostProcessor for RAG Generative AI notebook
- New Analyse Veterinary Documents with Healthcare NLP notebook
- Updated FewShot Assertion Classifier notebook
- New ALCOHOL SMOKING Demo
- New JSL vs GPT4 Demo
We Have Added and Updated a Substantial Number of New Clinical Models and Pipelines, Further Solidifying Our Offering in the Healthcare Domain.
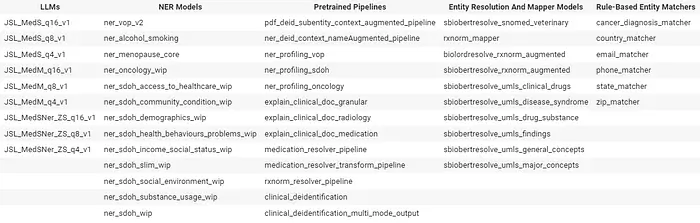
Trained And Updated Models-1 In v5.4
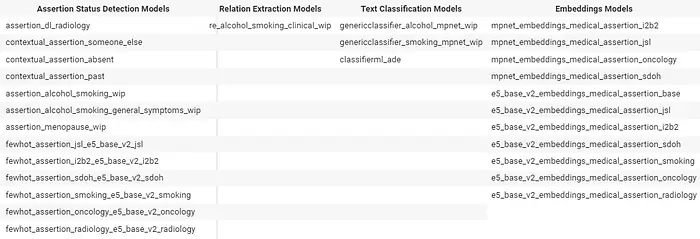
Trained And Updated Models-2 In v5.4
For all Spark NLP for Healthcare models, please check: Models Hub Page