John Snow Labs is thrilled to introduce a powerful set of new ONNX based clinical Named Entity Recognition (NER) models for English, Italian, and Spanish, in its’ most recent release expanding multilingual support beyond English for structured information extraction from clinical text. These models are designed to identify core medical entities including diseases, procedures, medications, and symptoms with high accuracy and production ready performance.
Building on our robust English foundation, these models expand our clinical NLP toolkit to support Italian and Spanish, making advanced entity extraction accessible for global teams.
Healthcare NLP
Healthcare NLP (Natural Language Processing) technologies extract insights from physician records, patient histories and diagnostic reports facilitating precise diagnosis.
Being the most widely used library in the healthcare industry, John Snow Labs’ Healthcare NLP comes with 2,500+ pretrained models that are all developed & trained with latest state-of-the-art algorithms to solve real world problems in the healthcare domain at scale. To provide reliable models and tools all the time while covering edge cases in real-world data and improve how well models generalize, the datasets and models are updated and augmented on a regular basis
Clinical NER in John Snow Labs
Named Entity Recognition (NER) is a Natural Language Processing (NLP) technique that involves identifying and extracting entities from a text, such as people, organizations, locations, dates, and other types of named entities.
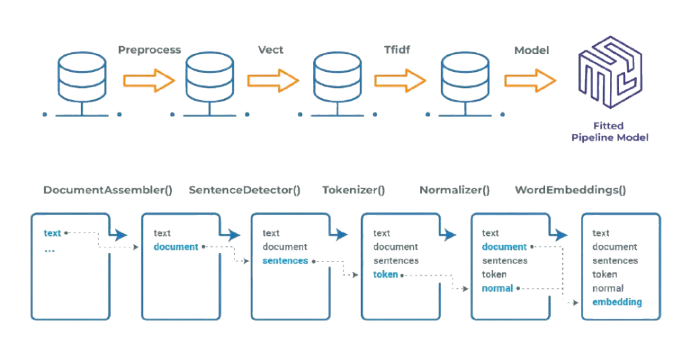
Spark NLP pipelines
Within the John Snow Labs ecosystem, clinical NER is implemented using transformer-based models integrated into scalable NLP pipelines. These pipelines handle document processing, tokenization, embeddings, and entity extraction in a modular way, making it possible to deploy NER consistently across batch and real time workloads.
For readers who want to explore how clinical NER works in practice without diving into model internals the following resource provides a practical overview with examples and pipelines:
Why ONNX?
ONNX (Open Neural Network Exchange) is a standardized format for representing machine learning models across frameworks and runtimes. In the context of clinical NLP, ONNX enables faster inference, reduced resource usage, and easier deployment across different environments, including cloud, on prem, and edge systems.
Available Models by Language
| Model Name | Language | Entities Extracted |
|---|---|---|
| roberta_med_ner_onnx | English | MEDICATION |
| roberta_disease_ner_onnx | English | DISEASE |
| roberta_procedure_ner_onnx | English | PROCEDURE |
| roberta_symptom_ner_onnx | English | SYMPTOM |
| bert_token_classifier_medical_ner_it_onnx | Italian | MEDICATION |
| bert_token_classifier_disease_ner_it_onnx | Italian | DISEASE |
| bert_token_classifier_procedure_ner_it_onnx | Italian | PROCEDURE |
| roberta_disease_ner_es_onnx | Spanish | DISEASE |
| roberta_procedure_ner_es_onnx | Spanish | PROCEDURE |
| roberta_symptom_ner_es_onnx | Spanish | SYMPTOM |
These models are built on transformer architectures such as BERT, RoBERTa, and XLM-RoBERTa, and are fine-tuned specifically for clinical and biomedical text. This allows them to handle domain specific terminology, abbreviations, and language specific clinical phrasing commonly found in real world medical narratives.
Multilingual Clinical Extraction
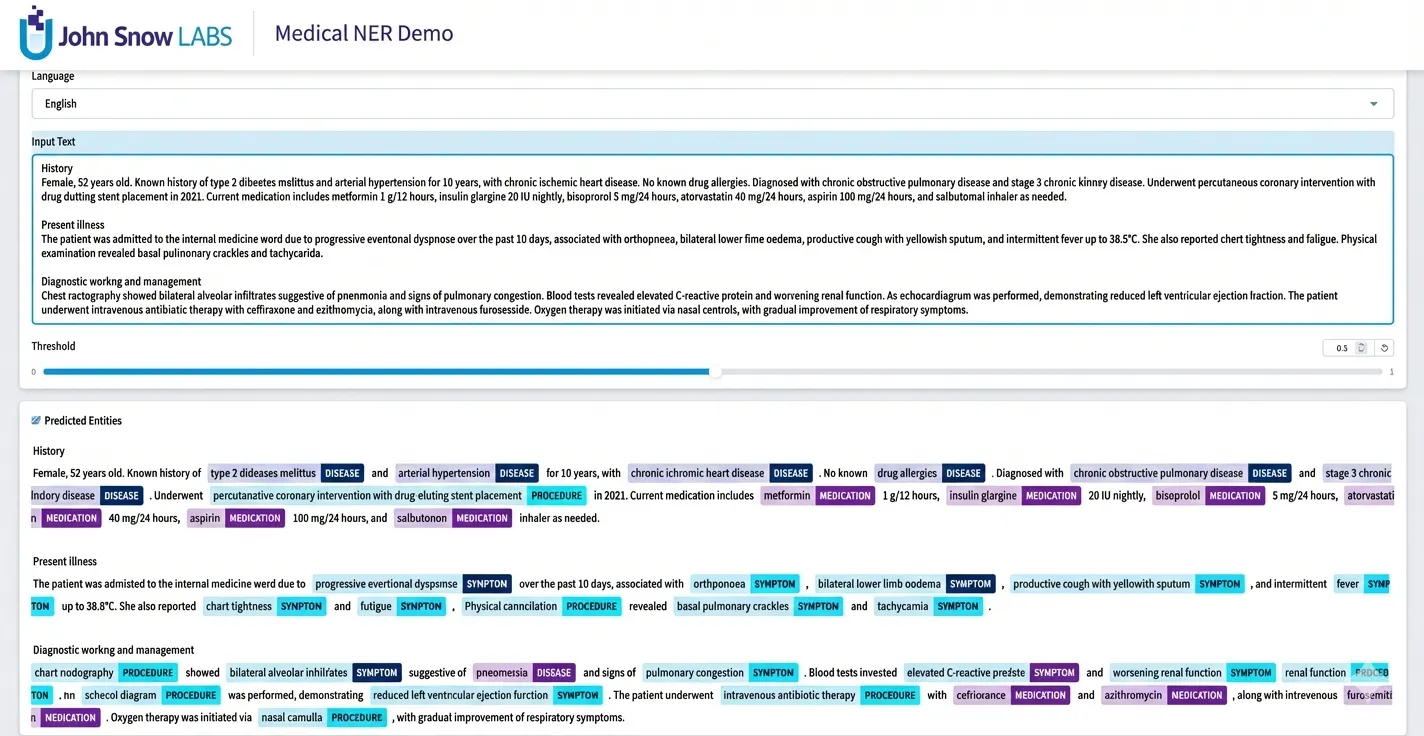
Clinical Named Entity Recognition powered by ONNX Runtime.
As demonstrated in the image above, the models are able to accurately parse and structure complex medical narratives across multiple categories.
Medical conditions such as “type 2 diabetes mellitus” and “arterial hypertension” are labeled as DISEASE, clinical interventions like “percutaneous coronary intervention” are recognized as PROCEDURE, and prescribed drugs including “metformin”, “insulin glargine”, and “aspirin” are identified as MEDICATION.
Usage examples
English Model
from johnsnowlabs import nlp, medical
# Start Spark session
spark = nlp.start()
# Build the pipeline
document_assembler = nlp.DocumentAssembler() \
.setInputCol("text") \
.setOutputCol("document")
tokenizer = nlp.Tokenizer() \
.setInputCols("document") \
.setOutputCol("token")
token_classifier = nlp.RoBertaForTokenClassification \
.pretrained("roberta_med_ner_onnx", "en", "clinical/models") \
.setInputCols(["document", "token"]) \
.setOutputCol("ner")
converter = nlp.NerConverter() \
.setInputCols(["document", "token", "ner"]) \
.setOutputCol("ner_chunk")
pipeline = nlp.Pipeline(stages=[
document_assembler, tokenizer,
token_classifier, converter
])
# Sample clinical text
sample_text = """The patient was diagnosed with type 2 diabetes mellitus and arterial hypertension.
She underwent percutaneous coronary intervention last year. Current medications include
metformin 500mg twice daily, insulin glargine 20 units at bedtime, and aspirin 81mg daily.
She reports occasional chest pain and shortness of breath."""
data = spark.createDataFrame([[sample_text]]).toDF("text")
result = pipeline.fit(data).transform(data)
Results:
result.selectExpr("explode(ner_chunk) as chunk").selectExpr(
"chunk.result as chunk",
"chunk.metadata['entity'] as entity"
).show(truncate=False)
+--------------------------------------+-----------+
| chunk | entity |
+--------------------------------------+-----------+
| metformin | MEDICATION|
| insulin glargine | MEDICATION|
| aspirin | MEDICATION|
+--------------------------------------+-----------+
Spanish Model
spanish_token_classifier = medical.MedicalBertForTokenClassifier \
.pretrained("bert_token_classifier_medical_ner_it_onnx", "it", "clinical/models") \
.setInputCols(["document", "token"]) \
.setOutputCol("ner")
pipeline = nlp.Pipeline(stages=[
document_assembler, tokenizer,
spanish_token_classifier, converter
])
sample_text_es = """El paciente fue diagnosticado con diabetes mellitus tipo 2 e hipertensión arterial.
Se le realizó una intervención coronaria percutánea el año pasado. Los medicamentos
actuales incluyen metformina 500mg dos veces al día, insulina glargina 20 unidades
al acostarse y aspirina 81mg diaria. Refiere dolor torácico ocasional y disnea."""
data_es = spark.createDataFrame([[sample_text_es]]).toDF("text")
result_es = pipeline_es.fit(data_es).transform(data_es)
Results:
result.selectExpr("explode(ner_chunk) as chunk").selectExpr(
"chunk.result as chunk",
"chunk.metadata['entity'] as entity"
).show(truncate=False)
+--------------------------------------+-----------+
| chunk | entity |
+--------------------------------------+-----------+
| metformina | MEDICATION|
| insulina glargina | MEDICATION|
| aspirina | MEDICATION|
+--------------------------------------+-----------+
Stay tuned for upcoming releases filling the noted gaps and expanding to more languages!
For additional information, please consult the following references:
- Healthcare NLP Release Notes: 6.3.0
- Documentation : NerDL, NerConverter, NerOverwriter.
- Python Docs : NerDL, NerConverter, NerOverwriter.
- Scala Docs : NerDL, NerConverter, NerOverwriter.
- For extended examples of usage, see the NerDL, NerConverter.
- For additional information, see Named Entity Recognition with BERT in Spark NLP, Text Chunking using Transformation-Based Learning.
Conclusion
The introduction of multilingual clinical NER models marks a significant milestone in John Snow Labs’ mission to globalize medical data extraction. By extending high-performing entity extraction capabilities across languages, these models bridge a critical gap in non-English clinical NLP, ensuring that healthcare providers and researchers worldwide can leverage structured insights from localized medical records.




























