How the John Snow Labs Terminology Server converts unstructured clinical narratives into standardized, machine-ready medical codes, bridging the gap between narrative medicine and healthcare analytics.
Every day, hospitals and health systems generate millions of clinical documents: discharge summaries, radiology reports, consultation notes, progress updates etc., that capture the full complexity of patient care. Yet the vast majority of that information remains locked in free text, invisible to the analytics systems, quality programs, and research platforms that depend on structured data.
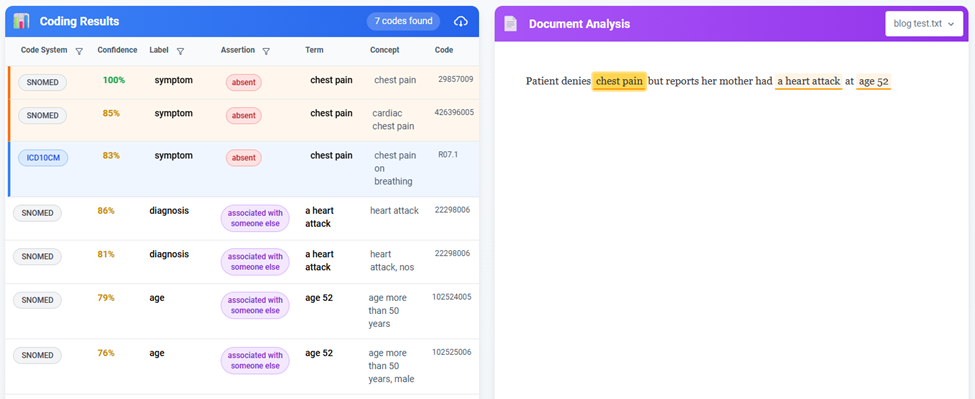
Consider this sentence from a real discharge summary: “Patient denies chest pain but reports her mother had a heart attack at age 52.” A clinician reading this immediately understands three distinct clinical facts: the patient does not currently have chest pain (negation), the patient carries a family history of myocardial infarction (relational context), and that event has a specific temporal anchor (age 52). To a conventional computer system, the same sentence is nothing more than a string of characters.
This gap between human-readable medical narrative and machine-processable healthcare data is one of the most consequential challenges in clinical informatics. An estimated 80% of all healthcare data lives in unstructured text. Closing that gap unlocks the ability to identify trial-eligible patients, automate quality measure calculation, support real-time clinical decisions, and reduce the crushing administrative burden on care teams.
John Snow Labs’ Terminology Server is built to close that gap.
Why 80% of Healthcare Data Is Clinically Invisible
Structured data in healthcare, such as lab values, vital signs or coded diagnoses, represents only a fraction of what is actually documented about a patient. The richest clinical intelligence lives in narrative text: the nuanced reasoning of a consulting physician, the detailed symptom chronology in a nurse’s progress note, the operative findings in a surgical report.
When that narrative remains unstructured, the downstream consequences are significant and measurable:
- Researchers cannot efficiently access real-world clinical data for studies, slowing evidence generation.
- Quality teams struggle to extract performance metrics from documentation, risking inaccurate reporting.
- Clinicians lack real-time decision support derived from their own patients’ longitudinal records.
- Administrators bear high costs for manual coding and chart abstraction that could be automated.
- Patients miss opportunities for clinical trials because their eligibility is buried in notes that are never computationally screened.
Traditional responses to this problem have relied on armies of medical coders manually reviewing documents and assigning standardized codes, a time-consuming, expensive, and error-prone process. Advances in medical natural language processing (NLP) and terminology science are fundamentally changing this equation.
Clinical Text Defies Simple Keyword Search: The Negation, Context, and Medical Nuance Challenge
Medical language is riddled with linguistic complexity that fundamentally alters clinical meaning. A naive keyword-matching system encountering the sentence “Patient denies chest pain” would flag “chest pain” as a finding and produce a clinically incorrect result. The same failure mode applies across the full range of documentation patterns that define everyday clinical writing:
- Negations: “No evidence of pneumonia”- the condition is explicitly absent.
- Historical references: “Prior history of diabetes”- relevant to context but not an active problem.
- Hypotheticals: “If symptoms worsen, consider admission”- a contingency, not a clinical event.
- Family history: “Mother with breast cancer”- the patient’s relative, not the patient.
- Temporal qualifiers: “Resolved fever”- a past finding that is no longer present.
Simply identifying medical terms is insufficient. Accurate clinical data extraction requires understanding what each term means in its specific linguistic and clinical context and the John Snow Labs Terminology Server is designed to do exactly that.
Clinical Document Intelligence for Automatic Coding of Unstructured Content
Imagine a system where you upload a clinical document and automatically receive:
✓ Every medical condition, procedure, and medication mentioned
✓ Clinical context for each (present, absent, historical, family history)
✓ Standardized codes from industry vocabularies (SNOMED CT, RxNorm, LOINC)
✓ OMOP Common Data Model mappings for research interoperability
✓ Structured data ready for EHR integration or analytics
This isn’t science fiction, it’s what modern clinical NLP combined with intelligent terminology services, can deliver.
What the John Snow Labs Terminology Server Is and How It Works
At its core, the Terminology Server is a medical concept normalization engine. Submit a phrase such as “heart attack,” “blood thinner,” or “sugar disease” and the server returns the authoritative, standardized code that represents that concept across every major clinical vocabulary. But the system’s capabilities extend well beyond simple lookup.
The Terminology Server combines two complementary retrieval strategies: string matching for exact and near-exact terms, and embedding-based semantic search for conceptually related expressions. This dual approach allows it to resolve synonyms, common misspellings, informal patient language, and clinical abbreviations without requiring the input text to precisely match a vocabulary entry.
“The fastest way to turn medical text into standardized codes, no matter the source.”
The platform comes pre-loaded with a widely used medical terminologies and offers a robust API and user interface that enables advanced concept search, mapping, and normalization. It addresses the challenges that frequently defeat traditional terminology servers: finding the right concept without an exact match, ranking results by clinical relevance, and mapping equivalent expressions across different vocabulary systems.
Comprehensive Medical Vocabulary Coverage: SNOMED CT, ICD-10, RxNorm, LOINC, and Beyond
The Terminology Server covers the full spectrum of clinical coding systems required for modern healthcare operations. Pre-loaded vocabularies include:
- SNOMED CT with 350,000+ clinical concepts spanning diagnoses, findings, procedures, and anatomy.
- ICD-10-CM/PCS, the international standard for diagnostic and procedure billing codes.
- RxNorm, the complete normalized medication vocabulary for drug names, formulations, and ingredients.
- LOINC covering 95,000+ laboratory observations and clinical measurements.
- CPT-4, Current Procedural Terminology for professional and outpatient services.
- MedDRA , the Medical Dictionary for Regulatory Activities, used in pharmacovigilance.
- ATC, the Anatomical Therapeutic Chemical classification for drug categorization.
- Additional vocabularies, NDC, HCPCS, ICD-9-CM, ICD-O3, MeSH, HPO, CVX, HGNC, and more.
This breadth of coverage means that clinical coders, researchers, and pharmacovigilance professionals can work within a single platform regardless of which vocabulary their specific workflow demands.
From Raw Clinical Text to Computable Data: The Three-Stage Processing Pipeline
The Terminology Server’s end-to-end workflow transforms unstructured clinical documents into standardized, analytics-ready data through three sequential stages: document ingestion, intelligent clinical NLP, and terminology resolution. Each stage is designed to handle the real-world complexity of clinical documentation without requiring manual preprocessing or configuration for standard use cases.
Stage 1 – Document Upload: Any Format, No Preprocessing Required
The pipeline begins with document submission. Clinical documents can be uploaded in any common format, PDF, Word, or plain text, without prior reformatting, cleaning, or structural transformation. This zero-friction ingestion is a deliberate design decision: real-world clinical documents arrive in heterogeneous formats, and requiring standardized input imposes an operational burden that undermines the value of automation.
Discharge summaries, radiology reports, consultation notes, operative reports, and progress updates can all be submitted directly, preserving the original content and formatting context that the NLP layer uses to interpret clinical meaning accurately.
Stage 2 – Intelligent Clinical NLP: Medical Entity Recognition with Context Awareness
The second stage is where clinical narrative becomes structured data. The system employs two complementary NLP approaches that work in concert: pre-built medical entity recognition models and custom zero-shot entity detection for specialized use cases.
Pre-Built Medical Entity Recognition
Trained medical NLP models automatically identify the full range of standard clinical entities present in the document:
- Diagnoses and conditions
- Procedures and interventions
- Medications and dosages
- Vital signs and measurements
Identifying these entities is necessary but not sufficient. The critical capability is that the system determines the clinical assertion status of every finding, the precise clinical meaning of what is being said, not just what terms appear:
- Present: the condition, finding, or event is currently true for this patient.
- Absent: the finding has been explicitly negated (“No evidence of pneumonia”).
- Historical: the finding relates to the patient’s past medical history, not the current encounter.
- Hypothetical: the finding is a future contingency, not an established clinical event.
- Family history: the finding applies to a relative of the patient, not to the patient directly.
This contextual assertion layer is what separates clinical NLP from keyword matching. “No evidence of diabetes” does not get coded as a diabetes diagnosis. “Mother with breast cancer” does not appear in the patient’s own condition list. The clinical meaning is preserved throughout the pipeline.
Zero-Shot Custom Entity Detection: Extracting Specialized Clinical Concepts Without Training Data
Standard medical NLP models are built around commonly documented clinical entities. But many of the most valuable use cases in healthcare involve entity types that are domain-specific, institution-specific, or research-specific, and for which no pre-trained model exists.
The Terminology Server addresses this through custom entity detection using zero-shot learning: the ability to define new entity types and extract them from clinical text without providing any labeled training examples. Organizations can deploy this capability immediately, without annotation projects or model retraining cycles.
Practical applications of zero-shot custom entity detection include:
- Research teams extracting “device complications” or “adverse events” as defined by their specific study protocol.
- Quality programs identifying “functional limitations” or “social determinants of health” from unstructured progress notes.
- Specialty clinics capturing domain-specific entities unique to their practice such as oncology response criteria, transplant-related findings, or rehabilitation outcomes.
This capability removes a historically significant barrier to clinical NLP adoption: the dependency on large annotated datasets for every new extraction task. Teams can define what they need and begin extracting it immediately.
Stage 3 – Terminology Resolution and Standardization: From Raw Text Spans to Computable Codes
The third stage transforms NLP output – raw text spans and their clinical assertions – into standardized, computable healthcare codes. A phrase like “heart attack” becomes SNOMED CT 22298006 (Myocardial infarction). A colloquial term like “blood thinner” resolves to its corresponding RxNorm entry. Every extracted entity is matched against the appropriate vocabulary and normalized to a canonical identifier.
The terminology resolution engine executes four operations on every extracted entity:
- Vocabulary resolution. Each entity is matched against the relevant medical vocabulary: SNOMED CT for conditions, RxNorm for medications, LOINC for laboratory observations, CPT for procedures, MedDRA for adverse events, and more.
- OMOP CDM mapping. Extracted entities can be mapped to OMOP standard concepts (leveraging vocabularies like SNOMED CT, RxNorm, and LOINC) when standard concept filters and vocabularies are applied, enabling downstream normalization into the OMOP Common Data Model for analytics and interoperability if needed.
- Semantic validation. The system confirms that mapped concepts are clinically coherent: medications map to drug concepts, laboratory tests map to measurement concepts, conditions map to condition concepts. Cross-domain mapping errors are caught at this stage.
- Clinical assertion preservation. The contextual status determined in Stage 2 is maintained through to the final coded output. A negated finding is coded as absent; a family history finding is coded with its relational qualifier. The structured output reflects clinical reality, not just terminology.
The result is a structured clinical dataset that is ready for EHR integration, research database population, quality reporting, or analytics, without any manual post-processing.
The Technology Behind Clinical Document Intelligence
Advanced NLP Models
The NLP layer of the Terminology Server is built on transformer architectures fine-tuned on millions of clinical documents. These models employ contextual embeddings that understand medical terminology in context rather than as isolated terms, and multi-task learning that simultaneously extracts entities and determines their clinical relationships. Zero-shot capabilities allow the system to generalize to new entity types without retraining, making it adaptable to emerging clinical documentation practices and research requirements.
Modern medical NLP leverages:
- Transformer architectures fine-tuned on millions of clinical documents
- Contextual embeddings that understand medical terminology in context
- Multi-task learning that simultaneously extracts entities and their relationships
- Zero-shot capabilities that work with custom entity types without retraining
OMOP Common Data Model
The Observational Medical Outcomes Partnership (OMOP) Common Data Model has become the standard architecture for large-scale real-world evidence generation and federated clinical research. The Terminology Server is fully aware of OMOP CDM conventions and can be constrained to return only OMOP Standard concepts, validating both concept designation and current concept validity in a single operation. The result is OMOP-ready data that integrates directly into research databases and federated data networks without additional transformation.
OMOP standardization enables:
- Multi-site research collaboration
- Cross-institution data exchange
- Interoperability with analytics platforms
- Integration with federated data networks
Enterprise-Grade Deployment: On-Premise, AWS, and Azure with HIPAA-Compliant Security
Healthcare data is among the most sensitive information an organization handles. The Terminology Server is designed with data sovereignty and regulatory compliance as foundational requirements, not afterthoughts.
Deployment Options
- On-Premise – install within your own secure network for maximum data control. All queries stay within the organization’s environment; no clinical text or patient data is transmitted over the public internet.
- AWS Marketplace – no-friction deployment on your virtual private cloud on Amazon Web Services.
- Azure Marketplace – seamless integration with your Azure environment for enterprise healthcare solutions.
Security and Compliance Architecture
- Encryption in transit and at rest
- Role-based access control for administrative functions
- Comprehensive audit logging of terminology updates and user access
- No persistent storage of patient identifiers or clinical notes – only standard terminologies and user-defined mappings
- Zero data sharing with John Snow Labs or any third party
- Compatibility with SAML/OAuth single sign-on when integrated via the UI
- MCP Server – Agent Connectivity and Tool Registration
The platform is engineered for high performance at enterprise scale, capable of serving sub-second responses under heavy workloads across large, combined vocabulary sizes. Its architecture supports horizontal scaling to handle high concurrency, and embedding-based search is accelerated with vector indexing to maintain speed as new terms are added. Batch operations leverage distributed processing for efficient large-dataset normalization. Vocabulary content is kept current through regular updates as new editions of ICD, SNOMED, LOINC, and other terminologies are released.
The platform provides step-by-step installation documentation and comprehensive API code samples to support rapid onboarding. Same business day 8×5 support is included with all subscriptions, and paying customers receive a private Slack channel for direct access to the John Snow Labs support team.
Real-World Clinical Applications: Four High-Impact Use Cases
Clinical Documentation Improvement and Medical Coding Automation
Hospital coding departments receive hundreds of discharge summaries daily. Instead of manually reviewing each document for billable diagnoses and procedures, coders receive auto-suggested codes extracted directly from the clinical narrative. The Terminology Server handles spelling variations, informal terminology, and synonym resolution automatically, capturing secondary diagnoses that manual review frequently overlooks and accelerating claim submission timelines.
Research Cohort Identification Across Clinical Vocabularies
Clinical research teams need to identify patient populations defined by precise, multi-criteria eligibility requirements – diagnoses, medications, exclusion conditions, and family history patterns – expressed differently across the notes of thousands of patients, providers, and time periods. Consider a cancer research team that needs to identify patients who have metastatic breast cancer, are currently on aromatase inhibitor therapy, do not have diabetes, and have no family history of cardiac disease. Executing this query against unstructured notes traditionally required weeks of manual chart review. With structured terminology output from the Terminology Server, the same query becomes a structured database operation.
Quality Measure Calculation and HEDIS Reporting
Health systems reporting HEDIS and other quality measures depend on structured clinical data: HbA1c test results, retinal and foot examination findings, blood pressure measurements, and documentation of specific interventions. The Terminology Server extracts and standardizes this information from clinical narratives, enabling data to flow directly into quality dashboards. Manual chart abstraction – one of the most resource-intensive activities in health system operations – is replaced by automated extraction that runs continuously as new documentation is created.
Prior Authorization and Administrative Burden Reduction
Insurance prior authorization requires extensive structured clinical documentation: diagnosis codes establishing medical necessity, records of previous treatments and their outcomes, current medication lists with dosages, and relevant laboratory values. The Terminology Server extracts and normalizes each of these elements from clinical narratives, enabling structured data to auto-populate prior authorization forms and reducing the administrative burden on clinical staff who would otherwise compile this documentation manually.
Target Users: Who Benefits Most from Medical Terminology Normalization
The Terminology Server is designed to serve a broad range of healthcare professionals and operational teams:
- Clinical Coders and Medical Billing Specialists – accurate code assignment from clinical notes, cross-mapping across ICD-10, SNOMED CT, and OMOP, and spelling/colloquialism handling for faster, more reliable coding.
- Researchers and Data Scientists – comprehensive cohort discovery across vocabularies, concept expansion for adverse event detection, and OMOP-ready output for federated research.
- Pharmacovigilance Professionals – MedDRA mapping from informal patient and clinical language, improved sensitivity for underreported events.
- Healthcare IT and Informatics Teams – robust API for workflow integration, enterprise security, flexible deployment, and regular vocabulary updates.
- Quality and Population Health Programs – automated metric extraction from clinical notes for HEDIS reporting, registry building, and population analytics.
The Future of Healthcare Analytics Is Making Existing Data Computable
The transformation from unstructured clinical narratives to standardized, computable healthcare data represents one of the most impactful advances available to healthcare IT today. By combining AI-powered medical NLP with comprehensive terminology services, health systems can:
- Reduce administrative burden on overtaxed clinical and coding staff
- Accelerate research with structured, analytics-ready real-world data
- Improve clinical decision support through better-organized patient information
- Enhance population health programs through comprehensive, structured analytics
- Lower operational costs by automating manual coding and chart abstraction
The technology is mature. The standards – SNOMED CT, ICD-10, RxNorm, LOINC, OMOP CDM – are well established. The regulatory environment supports it. The value case is documented across coding, research, quality, and pharmacovigilance applications.
The future of healthcare analytics isn’t about collecting more data – it’s about making the data we already have computable.
The question is no longer whether healthcare organizations will adopt medical terminology normalization, but how quickly they implement it to unlock the clinical and operational value hidden in their narratives.
Want to learn more about transforming your clinical documents into structured, standardized data? Explore John Snow Labs Terminology Service and see how automated medical entity extraction can accelerate your quality reporting, research initiatives, and clinical workflows.
Get John Snow Labs Terminology Server today: AWS, Azure , On Prem






























