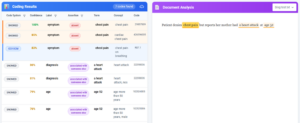
Clinical NLP extracts meaning from unstructured text. But in healthcare, extracted meaning isn’t useful until it speaks the same language as the systems that need to act on it.
An NLP model identifies “metformin 500mg” in a discharge summary. A medication reconciliation system needs NDC codes. An insurance claim needs HCPCS. A research database needs RxNorm. The entity is correct. But without standardization, it’s unusable by the downstream systems that depend on coded clinical data.
The following translation problem, from extracted entity to standard medical code, is where most clinical NLP workflows break down. Models get more accurate. Pipelines become fast. And then someone builds a manual coding step in between because there was no other option.
Generative AI Lab now addresses this directly with Medical Terminologies: a native capability that brings automated code resolution into annotation workflows, from pre-annotation through human review.
The standardization gap in clinical NLP
Healthcare operates on codes. ICD-10 for diagnoses. LOINC for lab observations. CPT for procedures. SNOMED CT for clinical concepts. RxNorm for medications. MeSH for biomedical literature and research. These aren’t just administrative conventions, they’re the shared language that makes clinical data interoperable across EHRs, billing systems, research databases, and regulatory submissions.
When NLP extracts entities from clinical text, those entities are surface forms: the words as they appear in the note. “Chest pain.” “Troponin I.” “Left knee arthroscopy.” Useful for reading. Not directly usable for coded data exchange.
Bridging that gap has traditionally required one of two things: a post-processing step that runs resolution after annotation, disconnected from the human review workflow, or manual coding by annotators who look up codes themselves, introducing inconsistency and slowing throughput considerably.
Neither approach scales. Neither keeps the coded result connected to the annotation decision that produced it. And neither supports the kind of Human-in-the-Loop workflow where a reviewer can verify both the entity label and the resolved code in the same interface.
Medical terminologies as a platform asset
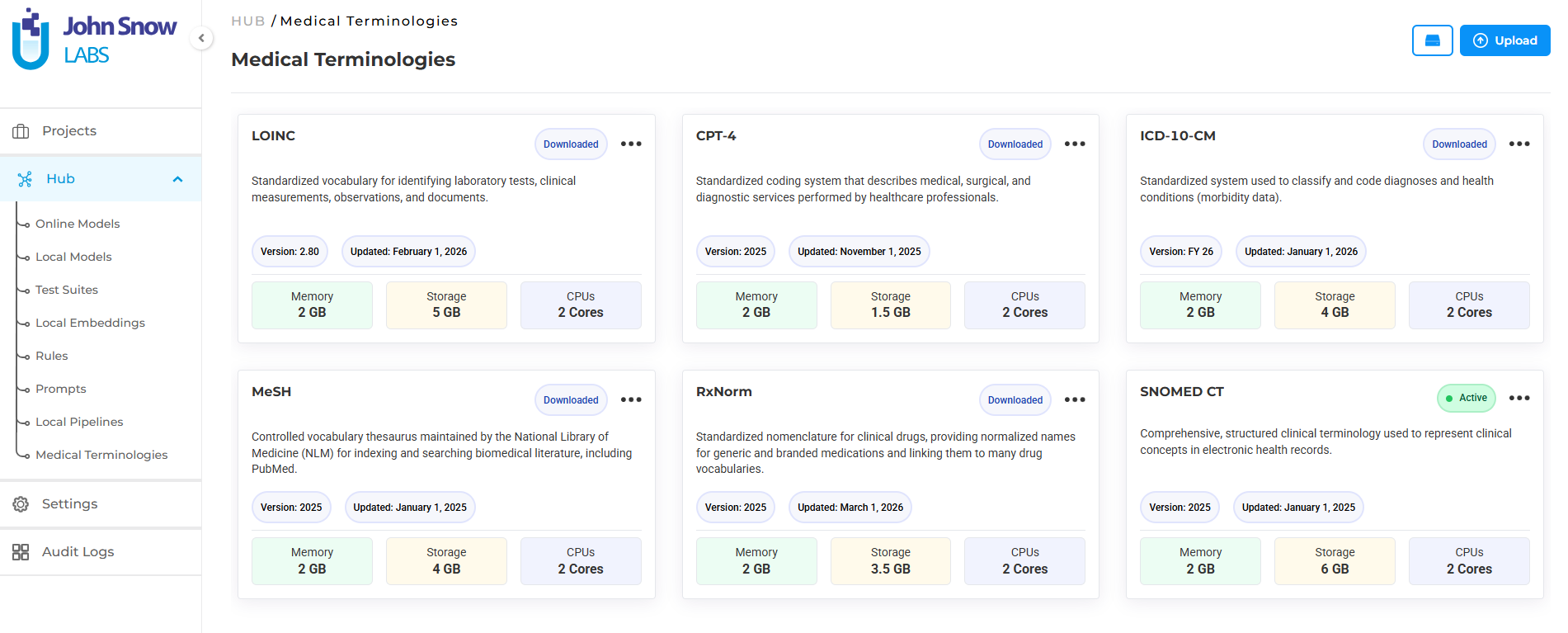
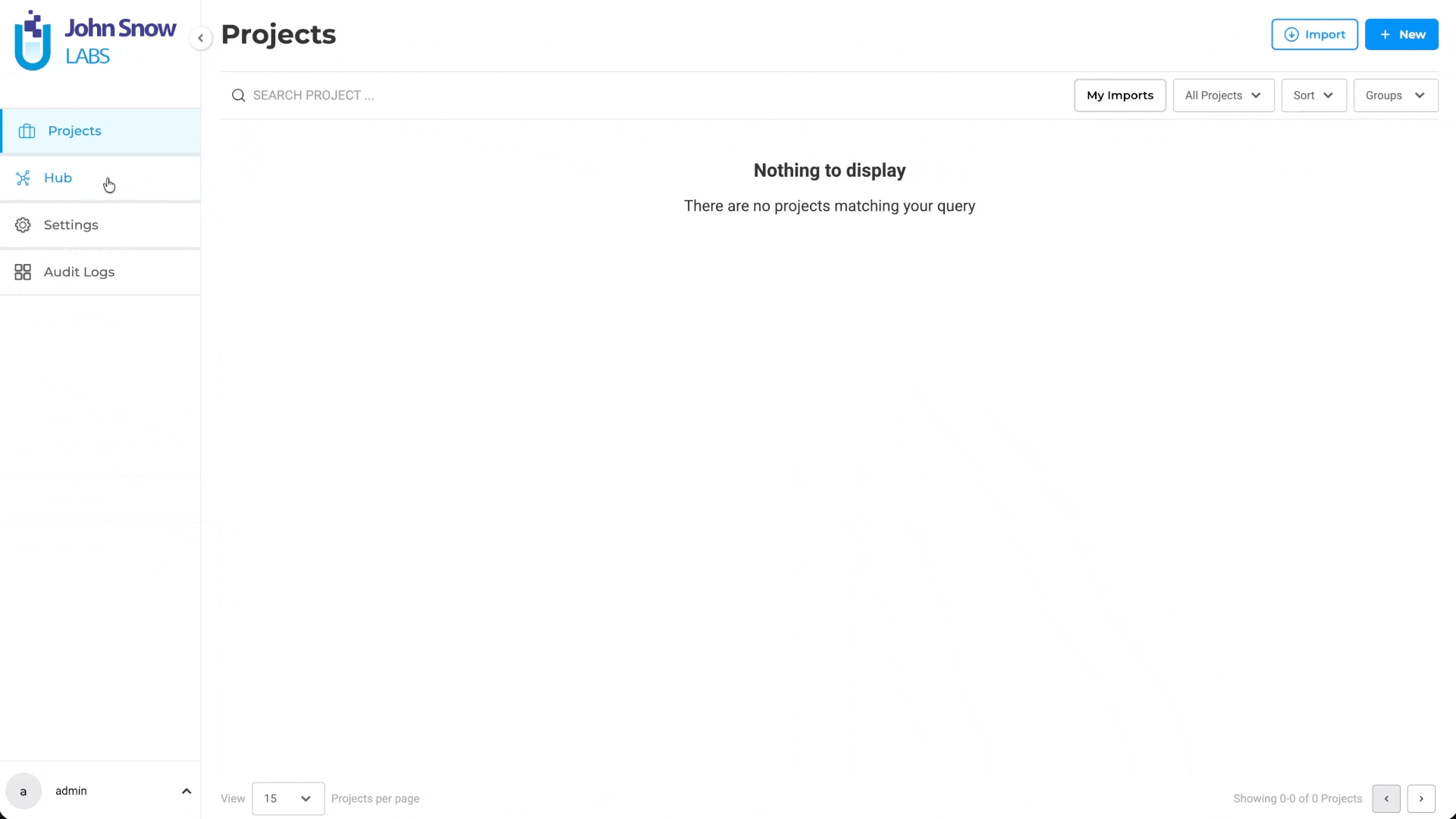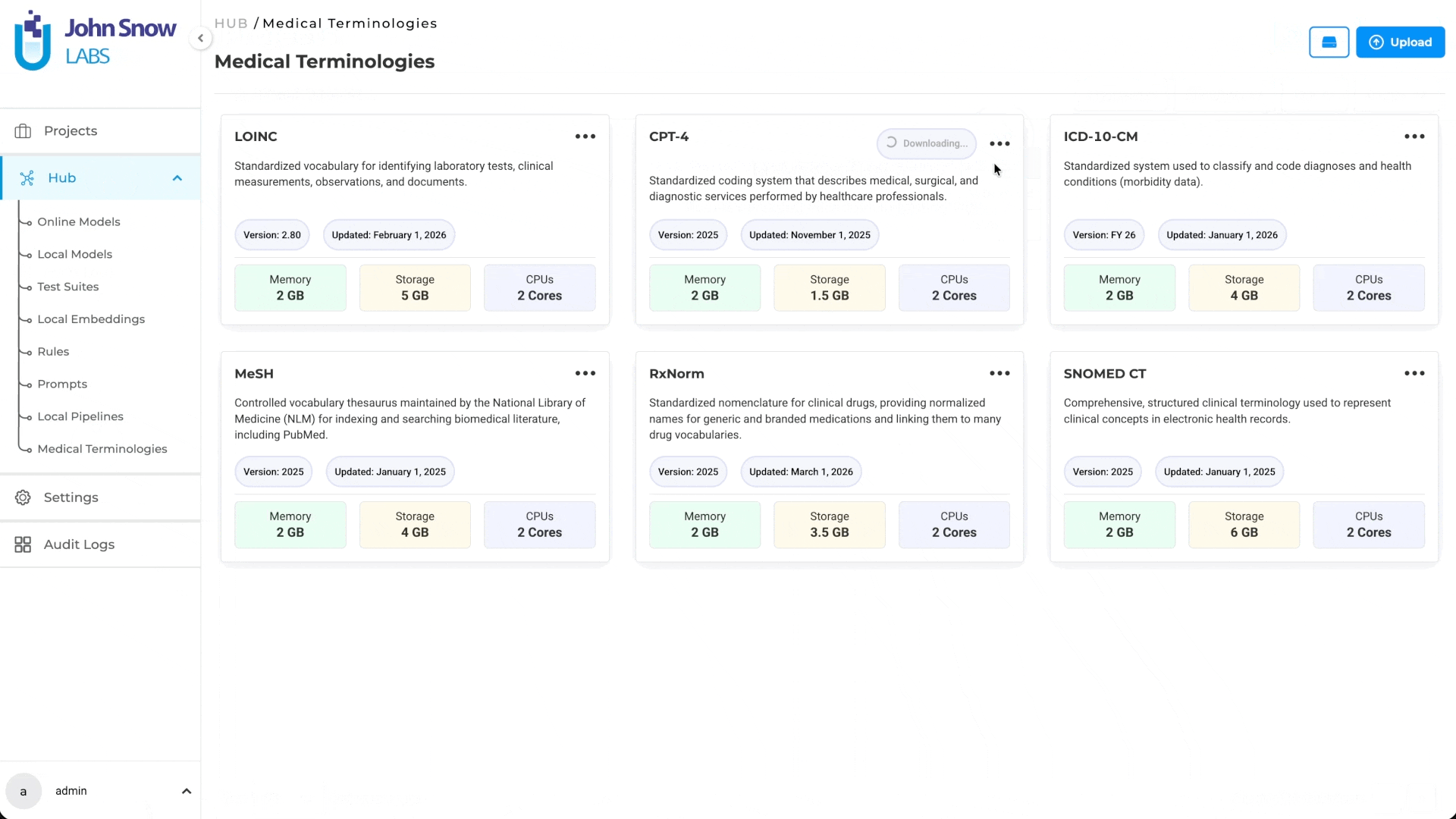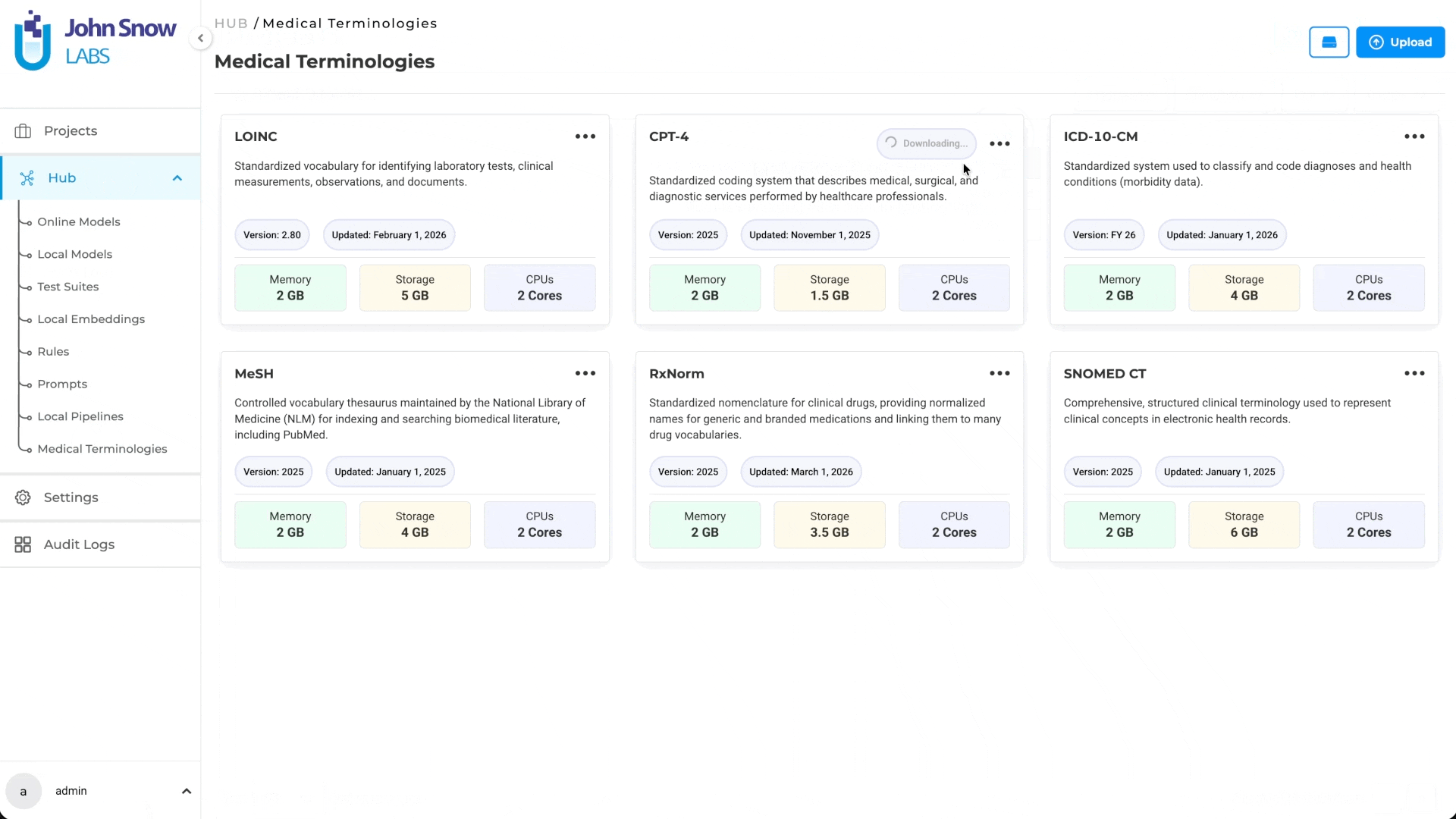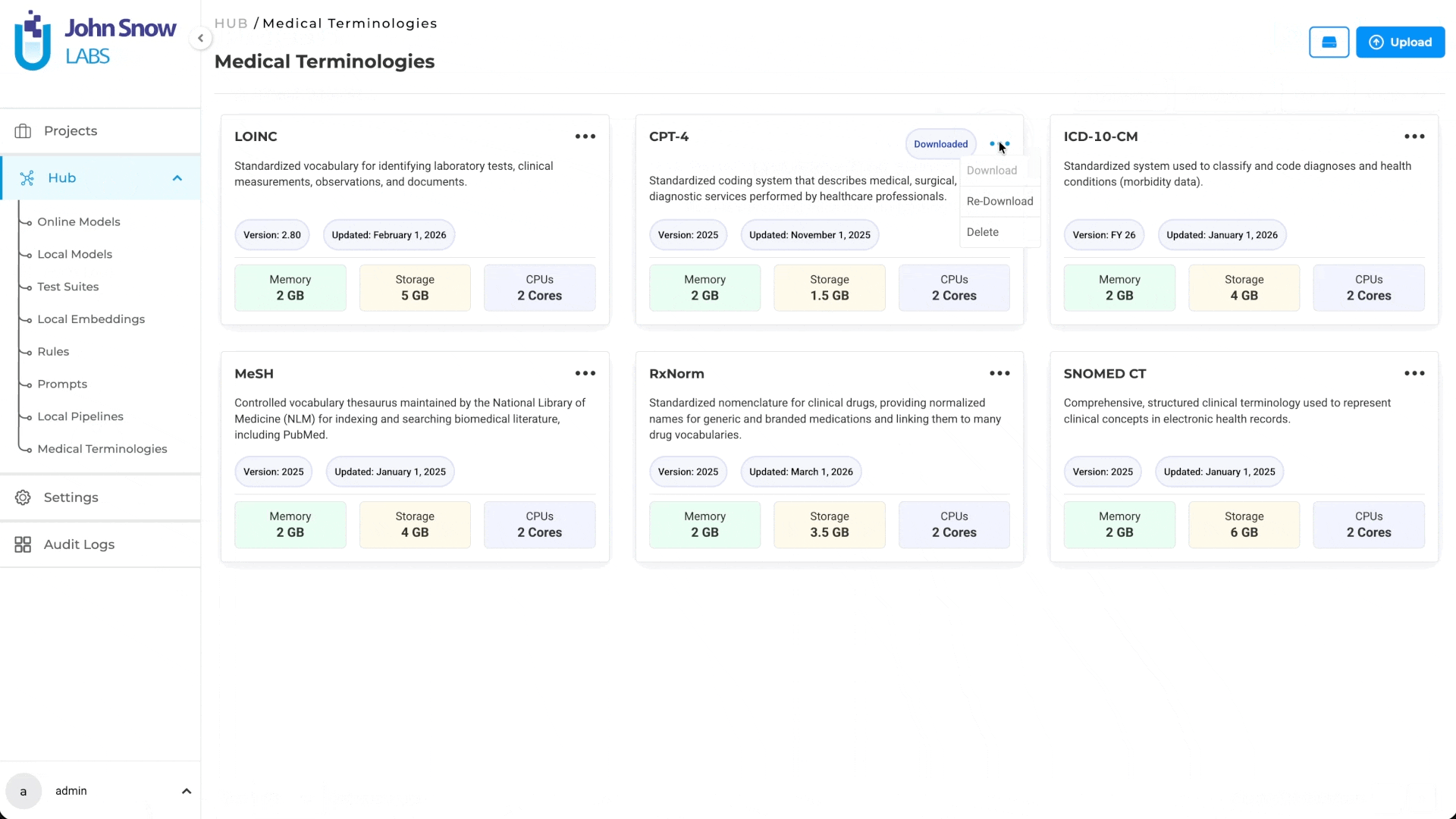
Medical Terminologies in Generative AI Lab are built as a first-class platform resource, not a post-processing plugin or an external lookup tool.
They live in the Hub alongside NLP models, where teams can browse available terminologies, understand their coverage, and download what their workflows require. Once downloaded, the terminology becomes a shared asset available across projects; teams working on different annotation initiatives can draw from the same resource without redundant setup.
Discovery follows the same licensing model as Healthcare NLP pipelines, which means terminology access is governed consistently with the rest of the platform’s clinical resources. Downloaded terminologies are available immediately for project configuration without additional infrastructure steps.
This matters because it changes how teams think about standardization. Instead of a separate system to maintain, medical code resolution becomes part of the same environment where annotation happens: discoverable, configurable, and deployable through the same workflow.
Connecting terminologies to annotation
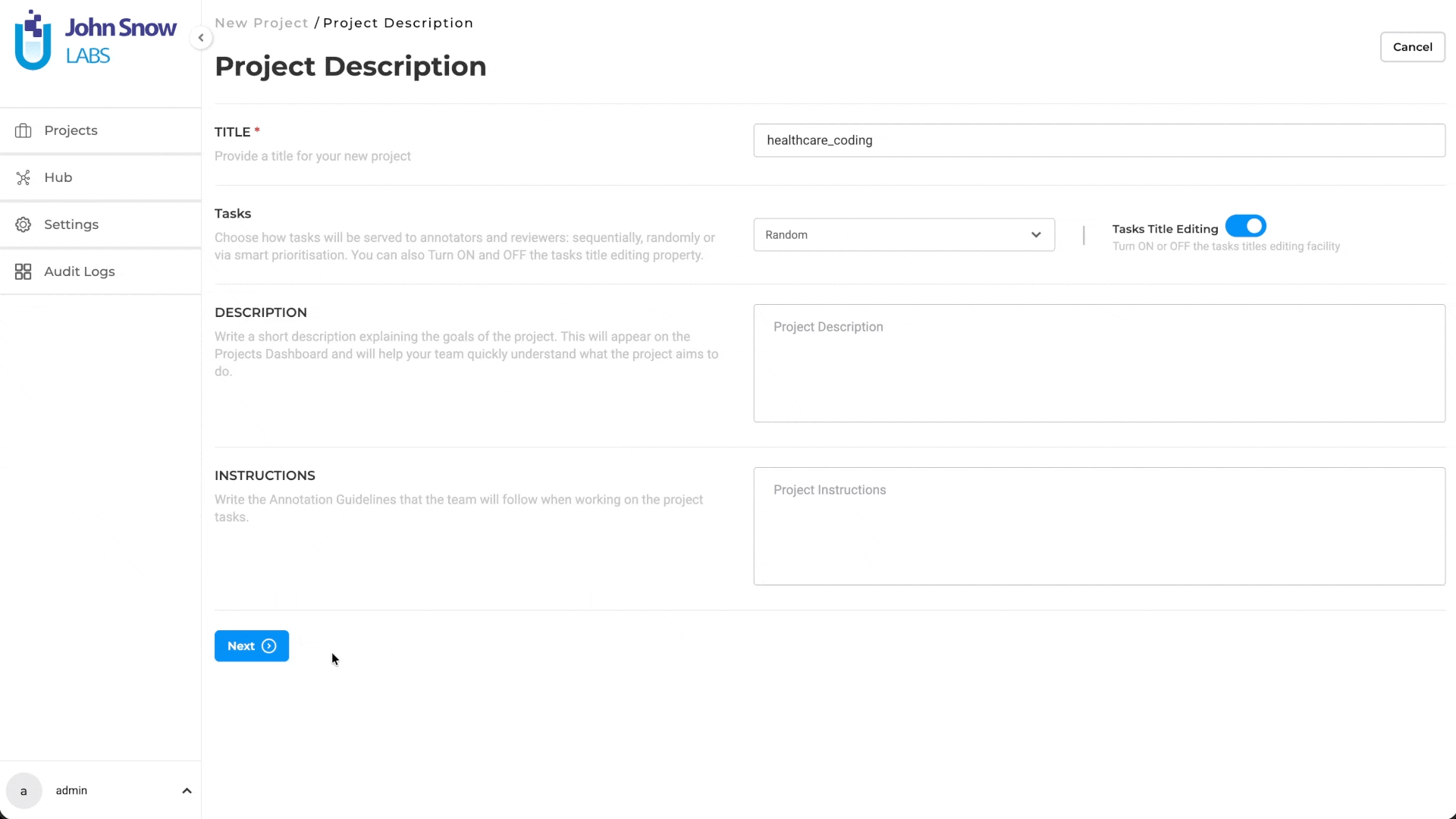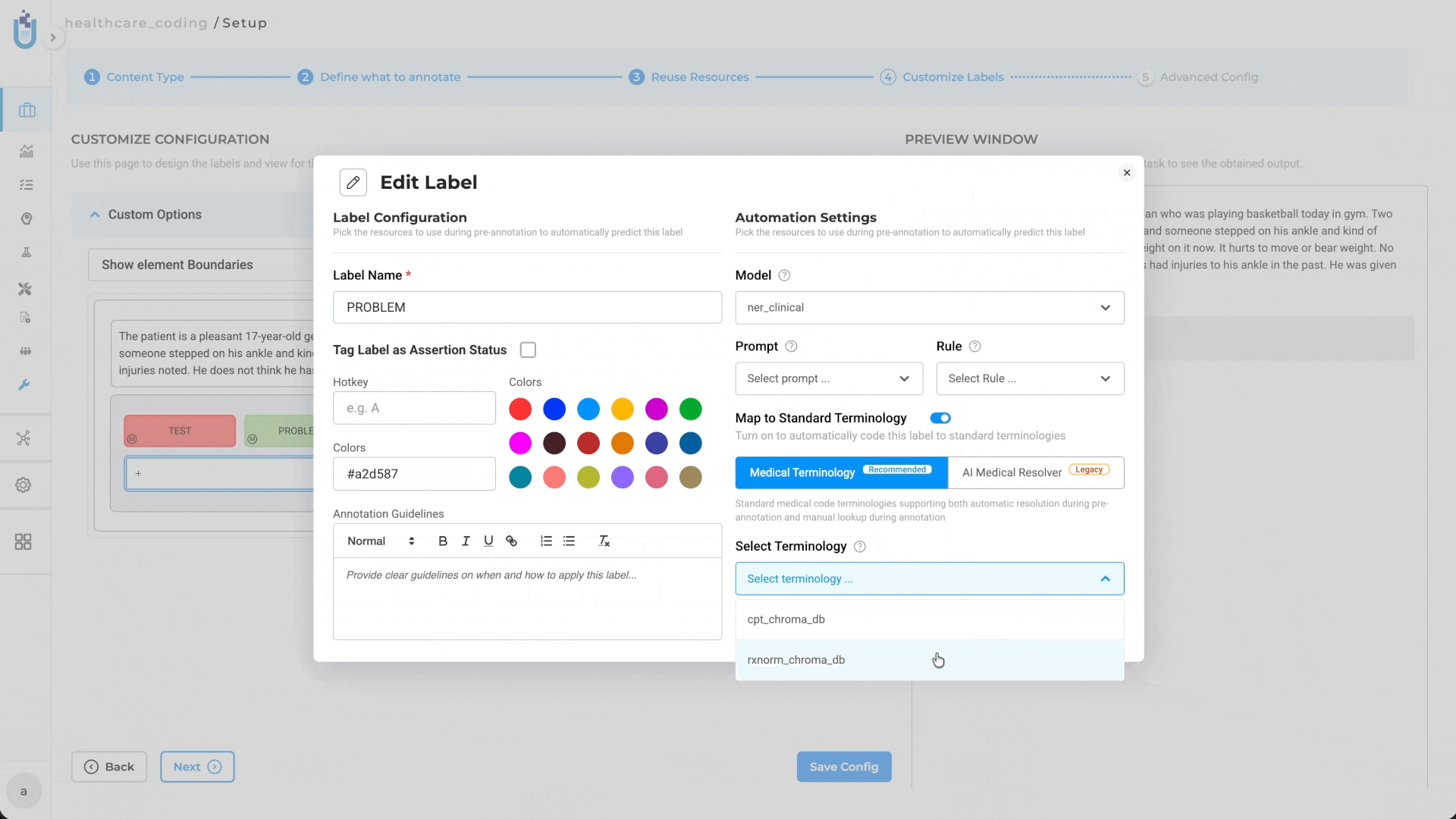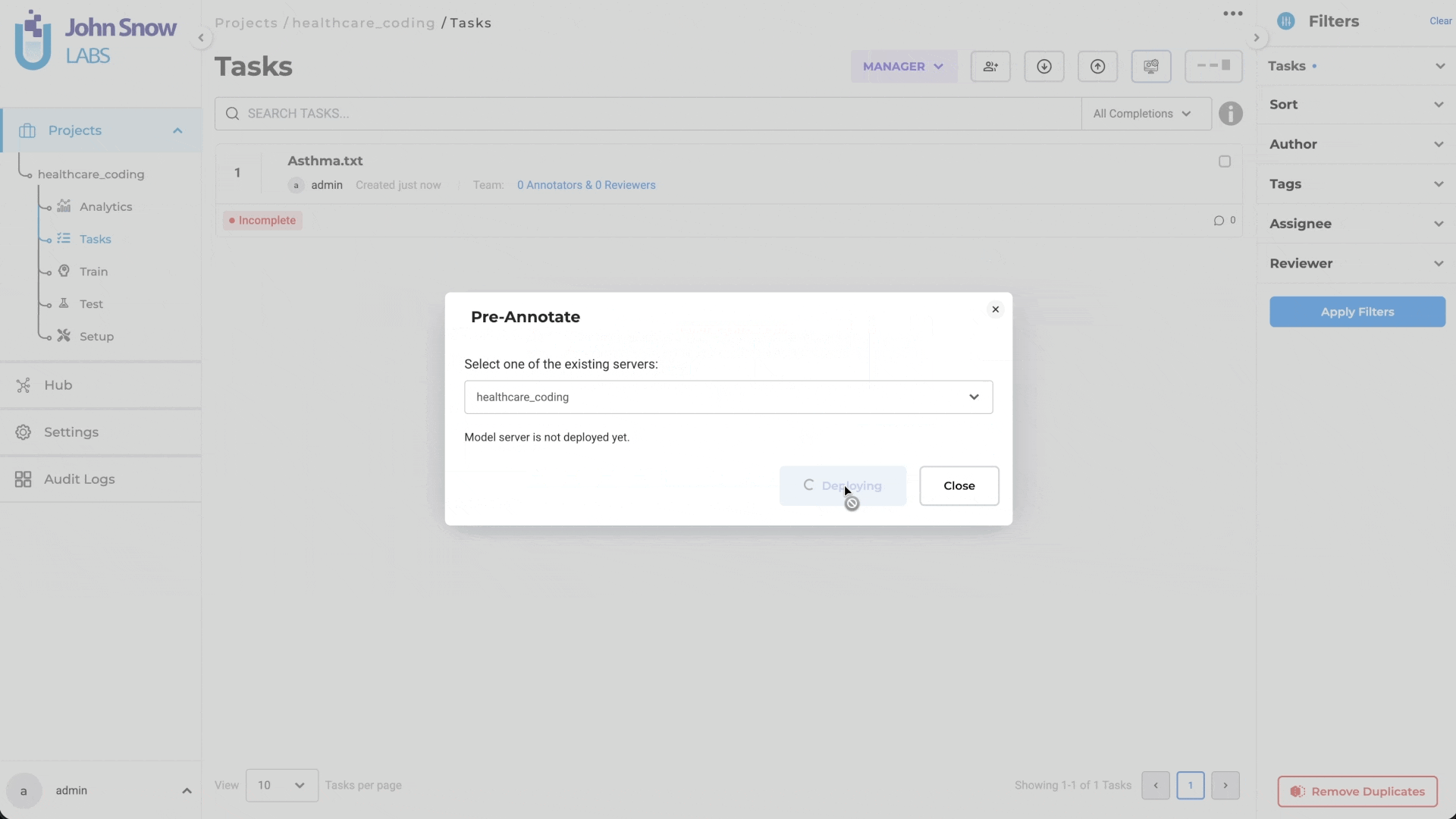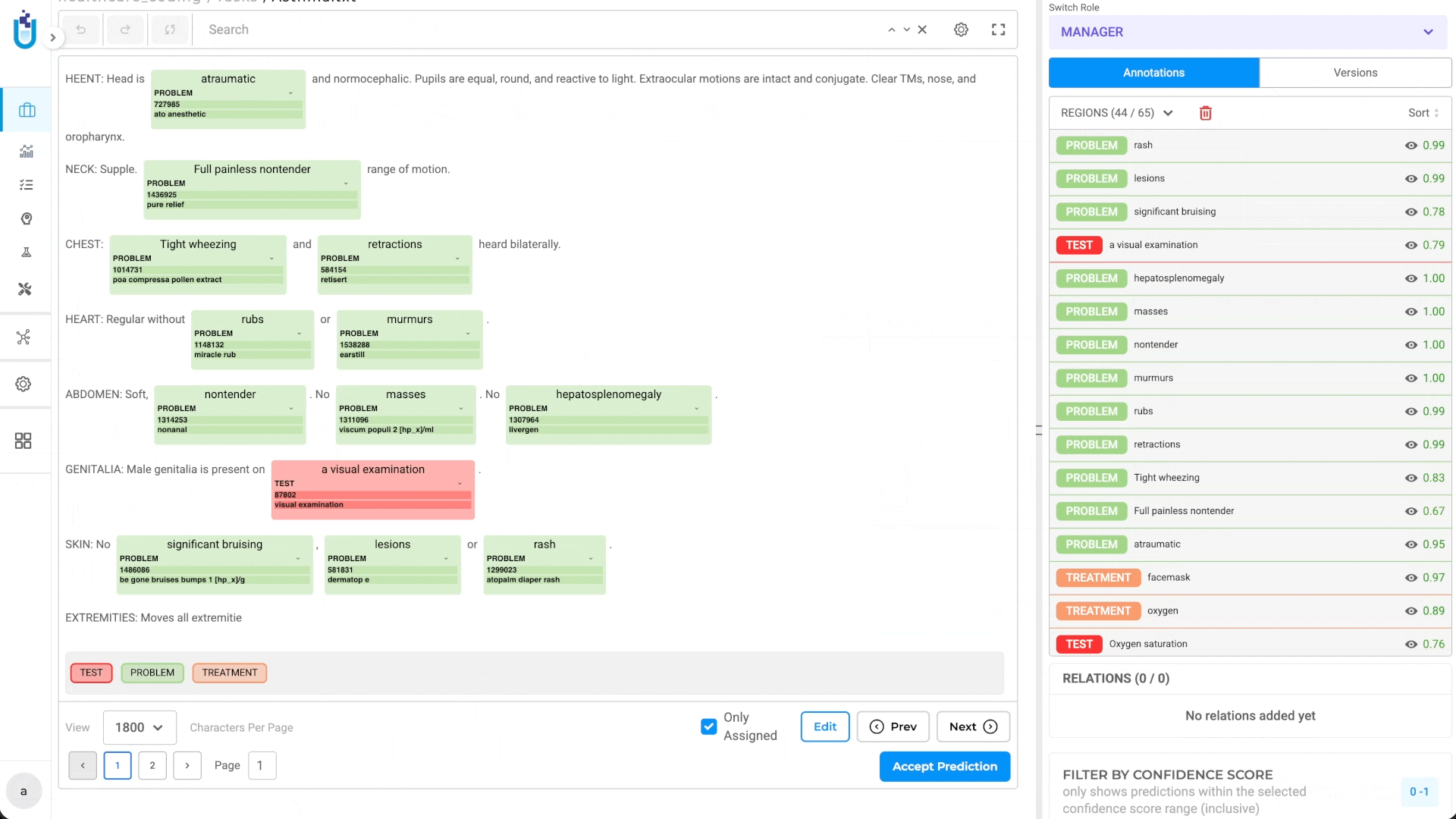
The integration point between medical terminologies and annotation work is the Label Configuration. When setting up a project, teams can link specific terminologies to specific labels: SNOMED CT to a diagnosis label, LOINC to a lab test label, RxNorm to a treatment label.
That linking isn’t just organizational. It determines how resolution happens during the two phases where it matters most.
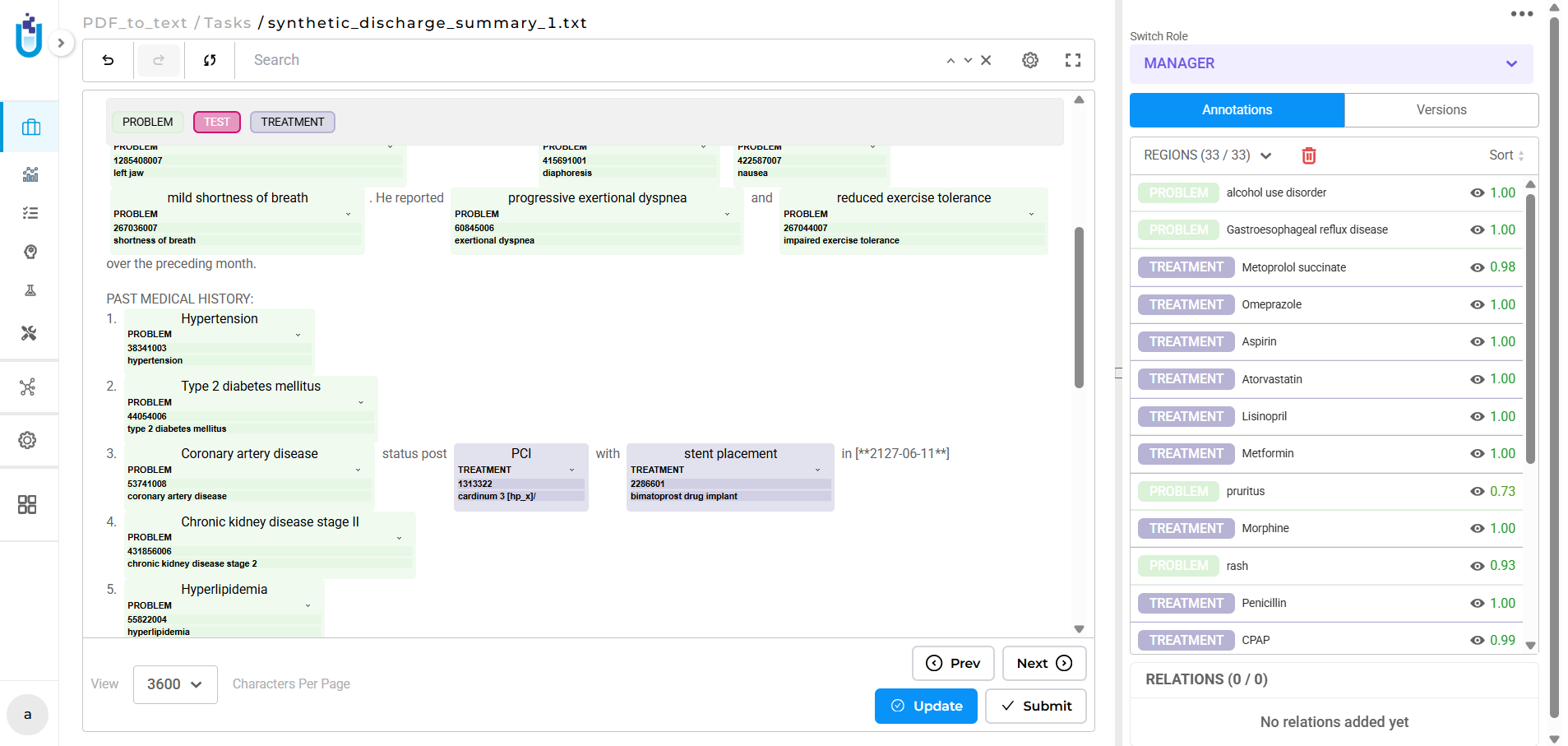
During pre-annotation, linked terminologies run automatically. When the pre-annotation pipeline processes a document and extracts an entity matching a configured label, the system resolves the corresponding medical code as part of the same pass. Annotators reviewing results see not just the entity span and label, but the resolved code that is ready to verify or correct, not to look up from scratch.
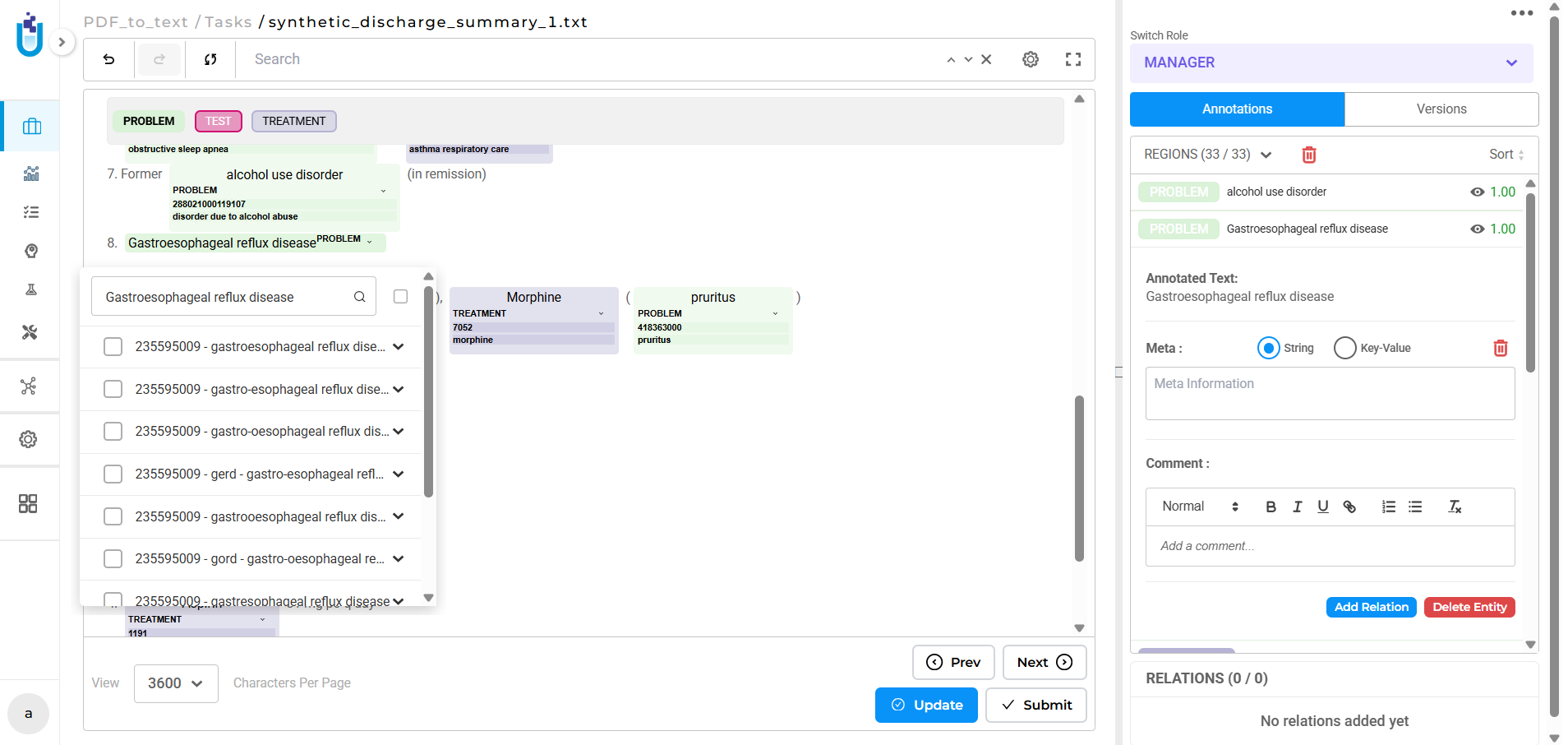
During human review, terminologies support manual lookup directly within the annotation interface. Annotators working on entities where automatic resolution didn’t produce a confident result can query the linked terminology interactively, selecting the correct code without leaving the platform.
Both modes keep the coded result attached to the annotation decision. The entity, the label, and the code travel together through the workflow and into the export.
Infrastructure that scales with complexity
Clinical annotation projects rarely need a single terminology. A project processing cardiology notes might require ICD-10 for diagnoses, LOINC for labs, and CPT for procedures simultaneously. Deploying and managing multiple terminologies across a team’s active projects would typically create significant infrastructure overhead.
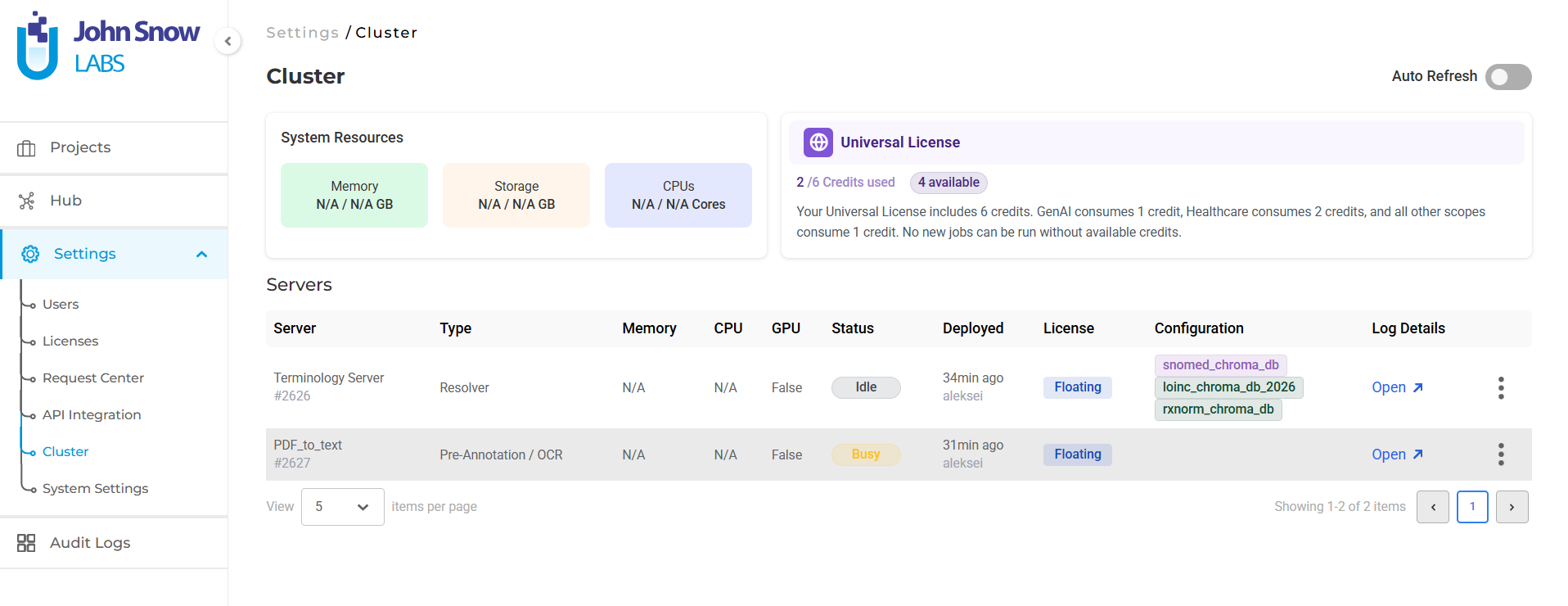
Generative AI Lab handles this through intelligent server management. When a project links to multiple terminologies, they deploy together on a single pre-annotation server rather than requiring separate instances for each. If a second project needs a subset of already-deployed terminologies, it reuses the active server rather than spinning up a new one. When a project needs a terminology not yet deployed, the system creates a new server containing the full required set.
The result is that teams configure what they need at the label level, and the platform handles deployment logistics. The Cluster page reflects what’s actually running: which terminologies are deployed on which server, uptime, and resource status giving administrators visibility without requiring manual coordination.
What this changes for clinical annotation workflows
The practical effect is that code resolution stops being a separate problem.
Annotation teams working on clinical documentation, discharge summaries, clinical trial records, prior authorization notes, structured intake forms, no longer need to treat entity extraction and code standardization as sequential steps managed by different tools. The terminology lookup happens where the annotation happens. The coded result is part of the annotation output.
For teams building training data for clinical NLP models, this means annotations carry richer supervision signal. Entity spans come with resolved codes, which means downstream tasks that require coded data, ICD prediction, lab value normalization, procedure coding, can draw on labeled examples that include both the surface form and the standard representation.
For compliance and interoperability work where coded accuracy matters as much as entity accuracy, it means human reviewers can verify both dimensions in a single pass. An annotator confirming a diagnosis label can simultaneously verify or correct the ICD-10 code without context switching.
The bigger picture
Healthcare AI’s value ultimately depends on whether its outputs connect to the systems that drive clinical decisions, billing, research, and regulatory compliance. Those systems speak in codes. The gap between what NLP extracts and what those systems consume has always required manual intervention to bridge.
Medical Terminologies in Generative AI Lab don’t eliminate the need for human judgment in that process; clinical coding is complex enough that human review remains essential. What they do is bring that judgment into the annotation workflow, where it can be applied consistently, at scale, with a complete record of the decisions made.
That’s what it takes to build clinical NLP pipelines that produce outputs the rest of the healthcare system can actually use.
Learn more
Full technical details on Medical Terminologies, Hub configuration, and deployment options:
👉 https://nlp.johnsnowlabs.com/docs/en/alab/release_notes
Questions about clinical NLP workflows, terminology configuration, or licensing? Reply to this email or reach out to our team.